Today I finally got a project out the door. BioStor is my take on what an interface to theBiodiversity Heritage Library (BHL) could look like. It features the visualisations I've mentioned in earlier posts, such as Google maps based on extracted localities, and tag trees. It also has a modified version of my earlier BHL viewer.
There are a number of ideas I want to play with using BioStor, but the main goal this site is to provide article-level metadata for BHL. As I've discussed earlier (see also Chris Freeland's post But where are the articles?), BHL has very little article-level metadata, making searching for articles a frustrating experience. BioStor aims to make this easier by providing an OpenURL resolver that tries to find articles in BHL.
BioStor supports the OpenURL standard, which means it can be used from
within EndNote and Zotero. Web sites that support COinS (such as Drupal-based Scratchpads and EOL's LifeDesks) can also be uses BioStor (see http://biostor.org/referrer.php for details).
My approach to finding articles in BHL is to take existing metadata from bilbiographies and databases, and use this to search BHL using techniques ranging from reasonably elegant (Smith-Waterman alignment on words to match titles) to down-and-dirty regular expression matching. Since this metadata may contain errors, BioStor provides basic editing tools (using reCAPTCHA rather than user logins at this point).
There's much to be done, the article finding is somewhat error-prone, and the search requires a local copy of BHL, and mine is rather out of date. However, it is a start.
To get a flavour of BioStor, try browsing some references:
http://biostor.org/reference/1
http://biostor.org/reference/4
http://biostor.org/reference/12
or view information for a journal:
http://biostor.org/issn/0007-1498
or an author:
http://biostor.org/author/41
http://biostor.org/author/16
or a taxon name:
http://biostor.org/name/Atelophryniscus%20chrysophorus
Rants, raves (and occasionally considered opinions) on phyloinformatics, taxonomy, and biodiversity informatics. For more ranty and less considered opinions, see my Twitter feed.
ISSN 2051-8188. Written content on this site is licensed under a Creative Commons Attribution 4.0 International license.
Monday, December 21, 2009
Friday, December 11, 2009
BHL interface ideas
I've been buried in programming (and it's exam time at Glasgow) so I've not blogged for a month (gasp). I've been playing with ways to visualise Biodiversity Heritage Library content for a while (click here for a list of previous posts), and have occasionally surfaced to tweet a screenshot via twitpic. The more I play with the BHL content the more I think it's a gold mine, and that many of the ideas I played with for my ill-fated Elsevier Challenge entry (website here, background paper at hdl:10101/npre.2009.3173.1) are taking on a new life with this project.
I'm hoping to release my BHL article finding and visualising web site by the end of the month, but meantime I'm gathering the screenshots here.
The first shows a Google map generated from latitude and longitudes extracted from OCR text using some simple regular expressions from page 7705952 in the BHL.There's quite a bit of latitude and longitude information in BHL, and that's before trying georeferencing tools.
The idea is to display this map next to the article so that user get's an immediate sense of what region in the world the article covers, such as this article about Riekia wasps:

I'm also interested in useful ways to display search results. Here's an experiment using TileBars to visualise how relevant a search result is. The width of the bar is a function of how many pages are in the article, the vertical stripes indicate pages that have the search term. The idea is to get a quick visual impression of whether the article mentions the term in parsing, or treats it in some detail.

TileBars were developed by Marti Hearst, whose web site has some great resources. Partly inspired by her BioText projec, as well as the thumbnail page display in JSTOR I'm now experimenting with showing thumbnails in search results. For example, here's a search for the deep sea octopus Graneledone pacifica, showing two articles:

I display thumbnails for pages that (a) have the name on the page, and (b) have what look like figure captions on them. The idea is that an article that figures a taxon is likely to be a fairly important article to look at, so displaying thumbnails will highlight those articles. The second article in the search results is the paper that published the name Graneledone pacifica, and the figures illustrate the taxon.
These are all pretty rough, but they give some idea of what I've been working on the last month.
I'm hoping to release my BHL article finding and visualising web site by the end of the month, but meantime I'm gathering the screenshots here.
The first shows a Google map generated from latitude and longitudes extracted from OCR text using some simple regular expressions from page 7705952 in the BHL.There's quite a bit of latitude and longitude information in BHL, and that's before trying georeferencing tools.
<

The idea is to display this map next to the article so that user get's an immediate sense of what region in the world the article covers, such as this article about Riekia wasps:

I'm also interested in useful ways to display search results. Here's an experiment using TileBars to visualise how relevant a search result is. The width of the bar is a function of how many pages are in the article, the vertical stripes indicate pages that have the search term. The idea is to get a quick visual impression of whether the article mentions the term in parsing, or treats it in some detail.

TileBars were developed by Marti Hearst, whose web site has some great resources. Partly inspired by her BioText projec, as well as the thumbnail page display in JSTOR I'm now experimenting with showing thumbnails in search results. For example, here's a search for the deep sea octopus Graneledone pacifica, showing two articles:

I display thumbnails for pages that (a) have the name on the page, and (b) have what look like figure captions on them. The idea is that an article that figures a taxon is likely to be a fairly important article to look at, so displaying thumbnails will highlight those articles. The second article in the search results is the paper that published the name Graneledone pacifica, and the figures illustrate the taxon.
These are all pretty rough, but they give some idea of what I've been working on the last month.
Labels:
BHL,
twitter,
visualisation
Wednesday, November 11, 2009
Tag trees: displaying the taxonomy of names in BHL
I've added a feature to my Biodiversity Heritage Library viewer that should help make sense of the names found on a page. Until now I've displayed them as a list of "tags", which ignores the relations among the names. Based on some code I'd developed for my e-Biosphere 09 challenge entry I've added a "tag tree" that displays the classification of the names found on a BHL page:

The idea is that a set of names can make much more sense if you know what kind of organism they are referring to. For example, I don't know what Onetes is, but if I look at BHL page 2298380 I can see that it's an insect:

The names in gray don't occur on the page, but do occur in the tree that links those names (the latter are highlighed in black). The tag tree can be useful for separating out host and parasite, e.g. BHL page 2298491 is about a flea and it's mammalian hosts:

The tag tree can also flag names that might be mistaken, such as those found on page 2298330:
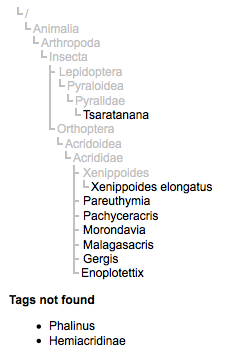

This page has names of some grasshoppers from Madagascar, as well as the name of a butterfly (Tsaratanana), which seems a little odd. Looking at the text, we discover that "Tsaratanana" is Mont. Tsaratanana a mountain in Madagascar. It would be fun to develop tools to annotate such cases so that somebody looking for the butterfly won't be presented with this page.
How it works
The inspiration for this tag tree came from several sources. David Remsen has often used an example of finding a fly name in the middle of a book on birds as being of interest, and the NCBI have a subtree view of taxa in a PubMed article. My own tag tree is constructed by finding for each name the ancestor-descendant path in a local, modified copy of the Catalogue of Life database, then assembling those paths into a tree. Because not all the names on a BHL page are in the Catalogue of Life, there may be names that aren't classified. These are simply listed below the tag tree (see image above).

The idea is that a set of names can make much more sense if you know what kind of organism they are referring to. For example, I don't know what Onetes is, but if I look at BHL page 2298380 I can see that it's an insect:

The names in gray don't occur on the page, but do occur in the tree that links those names (the latter are highlighed in black). The tag tree can be useful for separating out host and parasite, e.g. BHL page 2298491 is about a flea and it's mammalian hosts:

The tag tree can also flag names that might be mistaken, such as those found on page 2298330:

This page has names of some grasshoppers from Madagascar, as well as the name of a butterfly (Tsaratanana), which seems a little odd. Looking at the text, we discover that "Tsaratanana" is Mont. Tsaratanana a mountain in Madagascar. It would be fun to develop tools to annotate such cases so that somebody looking for the butterfly won't be presented with this page.
How it works
The inspiration for this tag tree came from several sources. David Remsen has often used an example of finding a fly name in the middle of a book on birds as being of interest, and the NCBI have a subtree view of taxa in a PubMed article. My own tag tree is constructed by finding for each name the ancestor-descendant path in a local, modified copy of the Catalogue of Life database, then assembling those paths into a tree. Because not all the names on a BHL page are in the Catalogue of Life, there may be names that aren't classified. These are simply listed below the tag tree (see image above).
Labels:
BHL,
Catalogue of Life,
tag tree,
tags
Monday, November 09, 2009
iTaxon screencast
Sadly I won't be at TDWG 2009, at least not in person. However, there is a session on wikis, which may contain this brief screencast of my iTaxon experiments. The screencast was made in haste, but tries to convey some of the ideas behind these experiments, especially the idea that by linking data together we can generate more interesting and rich views of objects such as scientific publications. The screencast starts with the The amphibian tree of life page.
Labels:
screencast,
TDWG,
wiki
Thursday, November 05, 2009
BHL Viewer now with go faster stripes
I'm using the Prototype Javascript library, and to my delight Bram Van Damme has written lazierLoad, inspired by Lazy Load for JQuery. lazierLoad works by attaching a listener to each image that listens for scroll events -- when the browser window scrolls each image receives a notification event and works out whether it needs to load the image. In theory, all you do is add the lazierLoad Javascript to your page, and only images that are currently visible will be fetched from the server. I say "in theory" because I needed to tweak the script a little because the thumbnails are inside a DIV element that has it's own scrollbar (thanks to the CSS style
overflow:auto). Hence I needed to add the listener to this DIV, and compute coordinates for the image taking the DIV into account. Like most things, easy once you know how (translation, after numerous failed attempts, and the occasional "doh!" it seems to work).You can see lazy loading in action if you view a BHL item, such as Item 26140. Note that this implementation of lazy loading doesn't work in Safari, much to my chagrin (it's my default browser). It works fine in Firefox
Labels:
BHL,
javascript,
lazy load
Tuesday, November 03, 2009
Biodiversity Heritage Library viewer experiments
In between the chaos that is term-time I've been playing with ways to view Biodiversity Heritage Library content. The viewer is crude, and likely to go off-line at any moment while I fuss with it, the you can view an example here. This link takes you to a display of BHL item 19513, which is volume 12 of the Bulletin of the British Museum (Natural History) Entomology, which includes some striking insects, such as the species of Lopheuthymia displayed below:

The viewer attempts to do several things:
Note that on the left hand side I'm displaying the articles that have already been found in the volume. The editing interface is crude, and I'll need to look at user authentication and versioning to do this seriously, but it seems a quick way to annotate BHL content. Much of this needs only be done once, as once we have article boundaries then searching BHL journal content using, say, OpenURL becomes easy, and we can link bibliographic records in nomenclators to BHL. Improving the metadata will also improve visualisation such as my BHL timeline.
I hope to play a bit more with the view over the next days and weeks. It's pretty simple (Javascript with PHP back end). The key to creating the viewer was a complete dump of BHL's page metadata kindly provided by Mike Lichtenberg. I use this to locate individual page images stored by the Internet Archive, which I then store locally (and generate 100 pixel wide thumbnails).
Potentially there's a lot more one could add to a tool like this. I'm playing with displaying the taxon names found by uBio so that I can flag instances where the page is where the name is first published. One could imagine adding other flags, such as when a taxon is depicted so that we could easily extract images of taxa from BHL content. It would also be nice to be able to add taxon names that uBio's algorithms have missed. Utlimately, one could even display the OCR text and correct/annotate that. Much to do...

The viewer attempts to do several things:
- Display a BHL document in a way that I can quickly scan pages looking for article boundaries by showing thumbnails
- Provide a simple way for me to edit metadata (if you find the start of an article you can click on the is the first page in an article and edit the article details)
- Provide a RIS dump of the articles which can then be uploaded into other bibliographic tools
Note that on the left hand side I'm displaying the articles that have already been found in the volume. The editing interface is crude, and I'll need to look at user authentication and versioning to do this seriously, but it seems a quick way to annotate BHL content. Much of this needs only be done once, as once we have article boundaries then searching BHL journal content using, say, OpenURL becomes easy, and we can link bibliographic records in nomenclators to BHL. Improving the metadata will also improve visualisation such as my BHL timeline.
I hope to play a bit more with the view over the next days and weeks. It's pretty simple (Javascript with PHP back end). The key to creating the viewer was a complete dump of BHL's page metadata kindly provided by Mike Lichtenberg. I use this to locate individual page images stored by the Internet Archive, which I then store locally (and generate 100 pixel wide thumbnails).
Potentially there's a lot more one could add to a tool like this. I'm playing with displaying the taxon names found by uBio so that I can flag instances where the page is where the name is first published. One could imagine adding other flags, such as when a taxon is depicted so that we could easily extract images of taxa from BHL content. It would also be nice to be able to add taxon names that uBio's algorithms have missed. Utlimately, one could even display the OCR text and correct/annotate that. Much to do...
Friday, October 23, 2009
n-gram fulltext indexing in MySQL
Continuing with my exploration of the Biodiversity Heritage Library one obstacle to linking BHL content with nomenclature databases is the lack of a consistent way to refer to the same bibliographic item (e.g., book or journal). For example, the Amphibia Species of the World (ASW) page for Gastrotheca aureomaculata gives the first reference for this name as:
The journal that ASW abbreviates as "Bull. U.S. Natl. Mus." is in the BHL, which gives its title as "Bulletin - United States National Museum.". How do I link these two records? In my bioGUID OpenURL project I've been doing things like using SQL LIKE statements with periods (.) replaced by wildcards ('%') to find journal titles that match abbreviations (as well as building a database of these abbreviations). But this is error prone, and won't work for abbreviations such as "Bull. U.S. Natl. Mus." because the word "National" has been abbreviated to "Natl", which isn't a substring of "National".
After exploring various methods (including longest common subsequences, and sequence alignment algorithms) I came across a MySQL plugin for n-grams. The plugin tokenises strings into bi-grams (tokens with just two characters, see the Wikipedia page on N-grams for more information). This means that even though as words "National" and "Natl" are different, they will have some similarity due to the shared bi-grams "Na" and "at".
So, I grabbed the source for the plugin and the ICU dependency, compiled the plugin and added it to MySQL (I'm running MySQL 5.1.34 on Mac OS X 10.5.8). The plugin can be added while the MySQL server is running using this SQL command:
Initial experiments seem promising. For the bhl_title table I created a bi-gram index:
If I then take the abbreviation "Bull. U.S. Natl. Mus.", strip out the punctuation, and search for the resulting string ("Bull US Natl Mus")
I get this:
The journal we want is the top hit (if only just). I'll probably have to do some post-processing to check that the top hit makes sense (e.g., is it a supersequence of the search term?) but this looks like a promising way to match abbreviated journal names and book titles to records in BHL (and other databases).
Gastrotheca aureomaculata Cochran and Goin, 1970, Bull. U.S. Natl. Mus., 288: 177. Holotype: FMNH 69701, by original designation. Type locality: "in [Departamento] Huila, Colombia, at San Antonio, a small village 25 kilometers west of San Agustín, at 2,300 meters".
The journal that ASW abbreviates as "Bull. U.S. Natl. Mus." is in the BHL, which gives its title as "Bulletin - United States National Museum.". How do I link these two records? In my bioGUID OpenURL project I've been doing things like using SQL LIKE statements with periods (.) replaced by wildcards ('%') to find journal titles that match abbreviations (as well as building a database of these abbreviations). But this is error prone, and won't work for abbreviations such as "Bull. U.S. Natl. Mus." because the word "National" has been abbreviated to "Natl", which isn't a substring of "National".
After exploring various methods (including longest common subsequences, and sequence alignment algorithms) I came across a MySQL plugin for n-grams. The plugin tokenises strings into bi-grams (tokens with just two characters, see the Wikipedia page on N-grams for more information). This means that even though as words "National" and "Natl" are different, they will have some similarity due to the shared bi-grams "Na" and "at".
So, I grabbed the source for the plugin and the ICU dependency, compiled the plugin and added it to MySQL (I'm running MySQL 5.1.34 on Mac OS X 10.5.8). The plugin can be added while the MySQL server is running using this SQL command:
INSTALL PLUGIN bigram SONAME 'libftbigram.so';
Initial experiments seem promising. For the bhl_title table I created a bi-gram index:
ALTER TABLE `bhl_title` ADD FULLTEXT (`ShortTitle`) WITH PARSER bigram;
If I then take the abbreviation "Bull. U.S. Natl. Mus.", strip out the punctuation, and search for the resulting string ("Bull US Natl Mus")
SELECT TitleID, ShortTitle, MATCH(ShortTitle) AGAINST('Bull U S Natl Museum')
AS score FROM bhl_title
WHERE MATCH(ShortTitle) AGAINST('Bull U S Natl Museum') LIMIT 5;
I get this:
| TitleID | ShortTitle | score |
|---|---|---|
| 7548 | Bulletin - United States National Museum. | 19.4019603729248 |
| 13855 | Bulletin du Muséum National d'Histoire Naturelle. | 17.6493873596191 |
| 14964 | Bulletin du Muséum National d'Histoire Naturelle. | 17.6493873596191 |
| 5943 | Bulletin du Muséum national d'histoire naturelle. | 17.6493873596191 |
| 12908 | Bulletin du Muséum National d'Histoire Naturelle. | 17.6493873596191 |
The journal we want is the top hit (if only just). I'll probably have to do some post-processing to check that the top hit makes sense (e.g., is it a supersequence of the search term?) but this looks like a promising way to match abbreviated journal names and book titles to records in BHL (and other databases).
Saturday, October 17, 2009
Memcached, Mac OS X, and PHP
Thinking about ways to improve the performance of some of my web servers I've begun to toy with Memcached. These notes are to remind me how to set it up (I'm using Mac OS X 10.5, Apache 2 and PHP 5.2.10, as provided by Apple). Erik's blog post Memcached with PHP on Mac OS X has a step-by-step guide, based on the post Setup a Memcached-Enabled MAMP Sandbox Environment by Nate Haug, and I've basically followed the steps they outline.
Now, I just need to explore how to actually use this...
- Install the Memcached service on Mac OS X: Follow the instructions in Nate Haug's post.
- Install Memcache PHP Extension: Apple's PHP doesn't come with the PECL package for memcache so download it. To compile it go:
phpize
./configure
make
sudo make install
One important point. If you are running 64-bit Mac OS X (as I am), ./configure by itself won't build a usable extension. However, a comment by Matt on Erik's original post provides the solution. Instead of just ./configure, type this at the command prompt:
MACOSX_DEPLOYMENT_TARGET=10.5 CFLAGS="-arch ppc -arch ppc64 -arch i386 -arch x86_64 -g -Os -pipe -no-cpp-precomp" CCFLAGS="-arch ppc -arch ppc64 -arch i386 -arch x86_64 -g -Os -pipe" CXXFLAGS="-arch ppc -arch ppc64 -arch i386 -arch x86_64 -g -Os -pipe" LDFLAGS="-arch ppc -arch ppc64 -arch i386 -arch x86_64 -bind_at_load" ./configure
Then follow the rest of Erik's instructions for adding the extension to your copy of PHP. - Restart Apache: You can do this by restarting Web sharing in System preferences. Use the phpinfo(); command to check that the extension is working. You should see something like this:
If you don't see this, something's gone wrong. The Apache web log may help (for example, that's where I discovered that I had the problem reported by several people who commented on Erik's post. - You can start the memcached daemon like this:
# /bin/sh
memcached -m 1 -l 127.0.0.1 -p 11211 -d
Now, I just need to explore how to actually use this...
Thursday, October 08, 2009
Linking Bulletin of Zoological Nomenclature to BHL

After some fussing and hair pulling I've constructed a demo of linking a journal to the Biodiversity Heritage Library and displaying the results in Zotero (see my earlier post for rationale).
After some searching I managed to retrieve metadata for several hundred article from the Bulletin of Zoological Nomenclature. Using a local copy of the BHL metadata, I wrote a script that looked up each article in BHL and found the URL of the first page in the article. I then created a Zotero group for this journal and uploaded the linked references.
You can browse the group, and if you belong to Zotero you can join the group and get a local copy of the references (which you can edit and correct, the harvesting won't be perfect).
I've also added these references to my bioGUID OpenURL resolver, making it easy to find a given article. For example, the OpenURL link http://bioguid.info/openurl/?genre=article&issn=0007-5167&volume=51&spage=7 displays a page for the article "Doris grandiflora Rapp, 1827 (currently Dendrodoris grandiflora; Mollusca, Gastropoda): proposed conservation of the specific name", together with a link to the article in BHL.
Wednesday, October 07, 2009
Zotero group for Biodiversity Heritage Library content
 One thing I find myself doing (probably more often than I should) is adding a reference to my Zotero library for an item in the Biodiversity Heritage Library (BHL). BHL doesn't have article-level metadata (see But where are the articles?), so when I discover a page of interest (e.g., one that contains the original description of a taxon) I store metadata for the article containing that page in my Zotero library. Typically this involves manually step back through scanned pages until I find the start of the article, then store that URL as well as the page number as a Zotero item. As an example, here is the record for Ogilby , J. Douglas (1907). A new tree frog from Brisbane. Proceedings of the Royal Society of Queensland 20:31-32. The URL in the Zotero record http://www.biodiversitylibrary.org/page/13861218 take you to the first page of this article.
One thing I find myself doing (probably more often than I should) is adding a reference to my Zotero library for an item in the Biodiversity Heritage Library (BHL). BHL doesn't have article-level metadata (see But where are the articles?), so when I discover a page of interest (e.g., one that contains the original description of a taxon) I store metadata for the article containing that page in my Zotero library. Typically this involves manually step back through scanned pages until I find the start of the article, then store that URL as well as the page number as a Zotero item. As an example, here is the record for Ogilby , J. Douglas (1907). A new tree frog from Brisbane. Proceedings of the Royal Society of Queensland 20:31-32. The URL in the Zotero record http://www.biodiversitylibrary.org/page/13861218 take you to the first page of this article.One reason for storing metadata in Zotero is so that these reference are made available through the bioGUID OpenURL resolver. This is achieved by regularly harvesting the RSS feed for my Zotero account, and adding items in that feed to the bioGUID database of articles. This makes Zotero highly attractive, as I don't have to write a graphical user interface to create bibliographic records. BHL have their own citation editor in the works ("CiteBank"), based on the ubiquitous Drupal, but I wonder whether Zotero is the better bet -- it has a bunch of nice features, including being able to sync local copies across multiple machines, and store local copies of PDFs (synced using WebDAV).
For fun I've created a Zotero group called Biodiversity Heritage Library which will list articles that I've extracted from BHL. At some point I may investigate automating this process of extracting articles (using existing blbiliographic metadata mapped to BHL page numbers), but for now there a mere 27 manually acquired items listed in the BHL group.
Tuesday, October 06, 2009
Wikipedia and Gregg's paradox
Continuing the theme of taxonomic classification in Wikipedia, I'm perversely delighted that Wikipedia demonstrates Gregg's paradox so nicely.
 The late John R. Gregg wrote several papers and a book exploring the logical structure of taxonomy. His 1954 book The language of taxonomy stimulated a debate a decade later in Systematic Zoology concerning what Buck and Hull (1966) (doi:10.2307/2411628) termed "Gregg's Paradox".
The late John R. Gregg wrote several papers and a book exploring the logical structure of taxonomy. His 1954 book The language of taxonomy stimulated a debate a decade later in Systematic Zoology concerning what Buck and Hull (1966) (doi:10.2307/2411628) termed "Gregg's Paradox".
Gregg showed that if we (a) treat taxa as sets defined by extension (i.e., by listing all members), and (b) accept that two sets with exactly the same content must be the same set, then many biological classifications violate these premises because the same taxon may be assigned to multiple levels in the Linnean hierarchy. For example, the aardvark, Orycteropus afer, is the only extant species of the genus Orycteropus, which is the only extant member of the family Orycteropodidae, which in turn is the sole extant representative of the order Tubulidentata. Under Gregg's model, Tubulidentata, Orycteropodidae, and Orycteropus are all the same thing as they have exactly the same content (i.e., Orycteropus afer). Put another way, monotypic taxa are redundant and violate basic set theory. Gregg would argue that they should be eliminated.
 Wikipedia illustrates this nicely. Wikipedia conforms to Gregg's model in that taxa are defined by extension (each taxon comprises one or more wiki pages), and if taxa have the same content only one taxon (typically that with the lowest taxonomic rank) has a page in Wikipedia. Put another way, if the aardvark is the sole representative of the Tubulidentata, then there is nothing that could be put on the Tubulidentata page that shouldn't also belong on the page for the aardvark. As a result, the page for the aardvark gives a full classification of this animal, but most taxa in the hierarchy don't have their own pages.
Wikipedia illustrates this nicely. Wikipedia conforms to Gregg's model in that taxa are defined by extension (each taxon comprises one or more wiki pages), and if taxa have the same content only one taxon (typically that with the lowest taxonomic rank) has a page in Wikipedia. Put another way, if the aardvark is the sole representative of the Tubulidentata, then there is nothing that could be put on the Tubulidentata page that shouldn't also belong on the page for the aardvark. As a result, the page for the aardvark gives a full classification of this animal, but most taxa in the hierarchy don't have their own pages.
Responses
There are several possible responses to Gregg's paradox. One is to argue that taxa should be defined intensionally (i.e., on the basis of their characters), which was Buck and Hull's approach. Essentially, they were arguing that we could (somewhat arbitrarily) specify properties of Orycteropodidae that weren't shared by all Tubulidentata, and hence we are justified in keeping these taxa separate. Gregg himself was less than impressed by this argument (doi:10.2307/2412017).
Another approach is to suggest that we may discover taxa in the future that will, say, be members of Orycteropus but which aren't O. afer, and that the taxa between the rank suborder and species are placeholders for these discoveries. Indeed, in the case of the Tubulidentata there are extinct aardvarks (doi:10.1163/002829675x00137, doi:10.1016/j.crpv.2005.12.016, and doi:10.1111/j.1096-3642.2008.00460.x) that could be added to Wikipedia, thus justifying the creation of pages for the taxa that Gregg would have us eliminate.
Of course, Gregg's paradox is a consequence of having ranks and requiring each rank (or at least a reasonable subset of them) to exist in a classification. If we ignore ranks, then there's no reason to put any taxa between Afrotheria and Orycteropus afer. So, we could drop this requirement for having taxa at each rank or, of course, drop ranks altogether, which is one of the motivations behind phylogenetic classifications (e.g., the phylocode).
Implications for parsing Wikipedia
From a practical point of view, Gregg's paradox means that one has to be careful parsing Wikipedia Taxoboxes. As I've argued earlier, the simplest way to ensure that a classification is a tree is for each taxon to include a unique parent taxon. The simplest way to extract this for a taxon in a Wikipedia page would be to retrieve the taxon immediately above it in the classification (i.e., for Orycteropus afer this would be Orycteropus). But Orycteropus doesn't have a page in Wikipedia (OK, it does, but it's a redirect to the page for the aardvark). So, we have to go up the classification until we hit Afrotheria before we get a taxon page.
Personally I quite like the fact that a largely forgotten argument from the middle of the last century concerning logic and Linnean taxonomy seems relevant again.
 The late John R. Gregg wrote several papers and a book exploring the logical structure of taxonomy. His 1954 book The language of taxonomy stimulated a debate a decade later in Systematic Zoology concerning what Buck and Hull (1966) (doi:10.2307/2411628) termed "Gregg's Paradox".
The late John R. Gregg wrote several papers and a book exploring the logical structure of taxonomy. His 1954 book The language of taxonomy stimulated a debate a decade later in Systematic Zoology concerning what Buck and Hull (1966) (doi:10.2307/2411628) termed "Gregg's Paradox". Gregg showed that if we (a) treat taxa as sets defined by extension (i.e., by listing all members), and (b) accept that two sets with exactly the same content must be the same set, then many biological classifications violate these premises because the same taxon may be assigned to multiple levels in the Linnean hierarchy. For example, the aardvark, Orycteropus afer, is the only extant species of the genus Orycteropus, which is the only extant member of the family Orycteropodidae, which in turn is the sole extant representative of the order Tubulidentata. Under Gregg's model, Tubulidentata, Orycteropodidae, and Orycteropus are all the same thing as they have exactly the same content (i.e., Orycteropus afer). Put another way, monotypic taxa are redundant and violate basic set theory. Gregg would argue that they should be eliminated.
 Wikipedia illustrates this nicely. Wikipedia conforms to Gregg's model in that taxa are defined by extension (each taxon comprises one or more wiki pages), and if taxa have the same content only one taxon (typically that with the lowest taxonomic rank) has a page in Wikipedia. Put another way, if the aardvark is the sole representative of the Tubulidentata, then there is nothing that could be put on the Tubulidentata page that shouldn't also belong on the page for the aardvark. As a result, the page for the aardvark gives a full classification of this animal, but most taxa in the hierarchy don't have their own pages.
Wikipedia illustrates this nicely. Wikipedia conforms to Gregg's model in that taxa are defined by extension (each taxon comprises one or more wiki pages), and if taxa have the same content only one taxon (typically that with the lowest taxonomic rank) has a page in Wikipedia. Put another way, if the aardvark is the sole representative of the Tubulidentata, then there is nothing that could be put on the Tubulidentata page that shouldn't also belong on the page for the aardvark. As a result, the page for the aardvark gives a full classification of this animal, but most taxa in the hierarchy don't have their own pages.Responses
There are several possible responses to Gregg's paradox. One is to argue that taxa should be defined intensionally (i.e., on the basis of their characters), which was Buck and Hull's approach. Essentially, they were arguing that we could (somewhat arbitrarily) specify properties of Orycteropodidae that weren't shared by all Tubulidentata, and hence we are justified in keeping these taxa separate. Gregg himself was less than impressed by this argument (doi:10.2307/2412017).
Another approach is to suggest that we may discover taxa in the future that will, say, be members of Orycteropus but which aren't O. afer, and that the taxa between the rank suborder and species are placeholders for these discoveries. Indeed, in the case of the Tubulidentata there are extinct aardvarks (doi:10.1163/002829675x00137, doi:10.1016/j.crpv.2005.12.016, and doi:10.1111/j.1096-3642.2008.00460.x) that could be added to Wikipedia, thus justifying the creation of pages for the taxa that Gregg would have us eliminate.
Of course, Gregg's paradox is a consequence of having ranks and requiring each rank (or at least a reasonable subset of them) to exist in a classification. If we ignore ranks, then there's no reason to put any taxa between Afrotheria and Orycteropus afer. So, we could drop this requirement for having taxa at each rank or, of course, drop ranks altogether, which is one of the motivations behind phylogenetic classifications (e.g., the phylocode).
Implications for parsing Wikipedia
From a practical point of view, Gregg's paradox means that one has to be careful parsing Wikipedia Taxoboxes. As I've argued earlier, the simplest way to ensure that a classification is a tree is for each taxon to include a unique parent taxon. The simplest way to extract this for a taxon in a Wikipedia page would be to retrieve the taxon immediately above it in the classification (i.e., for Orycteropus afer this would be Orycteropus). But Orycteropus doesn't have a page in Wikipedia (OK, it does, but it's a redirect to the page for the aardvark). So, we have to go up the classification until we hit Afrotheria before we get a taxon page.
Personally I quite like the fact that a largely forgotten argument from the middle of the last century concerning logic and Linnean taxonomy seems relevant again.
Monday, October 05, 2009
Wikipedia's taxonomic classification is badly broken
Wikipedia is wonderful, but parts of it are horribly broken. Take, for example, taxonomic classifications. A classification is a rooted tree, which means that each node in the tree has a single parent. We can store trees in databases in a variety of ways. For example, for each node we could store a list of its children, or we could store the single unique parent of each node. Ideally we'd choose to store one or other, but not both. If we store both sets of statements (i.e., that node A has node B as one of its children, and that node B's parent is node A) then there is enormous potential for these two statements to get out of sync.

This is what has happened in Wikipedia. Each page for a taxon lists the lineage to which it belongs (i.e., its parent, and its parent's parent, and so on), and also lists the children of that node. What this means is that if somebody edits the page for taxon A and adds taxon B as a child, they also need to edit the page for taxon B to make A its parent. If only one of these two edits is made the classification may end up internally inconsistent.
For example, the page for Amphibia lists the classification of Amphibia like this:
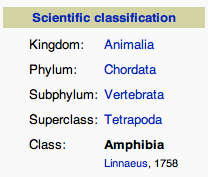
It also lists the child taxa of Amphibia:

So, the children of Amphibia are Temnospondyli, Lepospondyli, and Lissamphibia. Furthermore, Anura, Caudata, and Gymnophiona are children of Lissamphibia:

Given this, if I go to the pages for Anura, Caudata, and Gymnophiona I should see that each of these taxa lists Lissamphibia as its parent. However, only one of these (Caudata) does: the Anura and Gymnophiona both have Amphibia as their parents, not Lissamphibia.
The diagram below shows the taxa that have Amphibia as their parent:

Note that Stegocephalia have now turned up as an addition amphibian order, and that only Caudata is included in Lissamphibia. But what is striking is that another 274 Wikipedia taxon pages have Amphibia as their parent. These pages are all for fossil amphibians that do not fit easily in the existing Wikipedia classification.
From the perspective of building a database, the "has parent" relationship is the one I'd prefer to use, because that statement is going to be made just once (on the page for the taxon of interest). This seems a lot safer than making the statement "has child" on another page (for one thing, more than one page could claim a taxon as their child, which again will break the tree). But if we use the "has parent" relationship, our tree will be very bushy, with lots of fossil amphibian genera attached to the Amphibia node. This is going to make the tree hard to interpret, because this basal bush isn't saying that all these genera radiated off at once, but rather that we don't really know where in the amphibian tree these things go, so we'll have to settle for saying merely "they are amphibians" (for the cladistic theorists among you, this is Nelson and Platnick's "interpretation 2" in their "Multiple Branching in Cladograms: Two Interpretations", doi:10.2307/2412630).
So, the dilemma is whether to use "has child" relationships, and accept that these are likely to be inconsistent with the inverse "has parent" relationship, or use the "has parent" relationship, which will be internally consistent, but at the cost of potentially very large, unresolved bushes due to fossil taxa of uncertain affinities.

This is what has happened in Wikipedia. Each page for a taxon lists the lineage to which it belongs (i.e., its parent, and its parent's parent, and so on), and also lists the children of that node. What this means is that if somebody edits the page for taxon A and adds taxon B as a child, they also need to edit the page for taxon B to make A its parent. If only one of these two edits is made the classification may end up internally inconsistent.
For example, the page for Amphibia lists the classification of Amphibia like this:

It also lists the child taxa of Amphibia:

So, the children of Amphibia are Temnospondyli, Lepospondyli, and Lissamphibia. Furthermore, Anura, Caudata, and Gymnophiona are children of Lissamphibia:

Given this, if I go to the pages for Anura, Caudata, and Gymnophiona I should see that each of these taxa lists Lissamphibia as its parent. However, only one of these (Caudata) does: the Anura and Gymnophiona both have Amphibia as their parents, not Lissamphibia.
The diagram below shows the taxa that have Amphibia as their parent:

Note that Stegocephalia have now turned up as an addition amphibian order, and that only Caudata is included in Lissamphibia. But what is striking is that another 274 Wikipedia taxon pages have Amphibia as their parent. These pages are all for fossil amphibians that do not fit easily in the existing Wikipedia classification.
From the perspective of building a database, the "has parent" relationship is the one I'd prefer to use, because that statement is going to be made just once (on the page for the taxon of interest). This seems a lot safer than making the statement "has child" on another page (for one thing, more than one page could claim a taxon as their child, which again will break the tree). But if we use the "has parent" relationship, our tree will be very bushy, with lots of fossil amphibian genera attached to the Amphibia node. This is going to make the tree hard to interpret, because this basal bush isn't saying that all these genera radiated off at once, but rather that we don't really know where in the amphibian tree these things go, so we'll have to settle for saying merely "they are amphibians" (for the cladistic theorists among you, this is Nelson and Platnick's "interpretation 2" in their "Multiple Branching in Cladograms: Two Interpretations", doi:10.2307/2412630).
So, the dilemma is whether to use "has child" relationships, and accept that these are likely to be inconsistent with the inverse "has parent" relationship, or use the "has parent" relationship, which will be internally consistent, but at the cost of potentially very large, unresolved bushes due to fossil taxa of uncertain affinities.
Friday, October 02, 2009
Biodiversity Heritage Library sparklines
Time for a quick and dirty Friday afternoon hack. Based on responses to the BHL timeline I released two days ago, I've created a version that can compare the history of two names using sparklines (created using Google's Chart API). I use sparklines to give a quick summary of hits over time (grouped by decade).
The demo is here. It's crude (minimal error checking, no progress bars while it talks to BHL), but it's home time. As an example, here is a screen shot comparing the occurrences in BHL for two rival names for the sperm whale, Physeter catodon and Physeter macrocephalus:

There is a link to the full timeline for each of these names so you can investigate more. Note that the sparklines will be heavily biased by BHL coverage, but it may yield some interesting insights into the history of the usage of a name.
The demo is here. It's crude (minimal error checking, no progress bars while it talks to BHL), but it's home time. As an example, here is a screen shot comparing the occurrences in BHL for two rival names for the sperm whale, Physeter catodon and Physeter macrocephalus:

There is a link to the full timeline for each of these names so you can investigate more. Note that the sparklines will be heavily biased by BHL coverage, but it may yield some interesting insights into the history of the usage of a name.
Labels:
BHL,
sparklines,
timeline,
visualisation
Thursday, October 01, 2009
Co-located Collaborative Tree Comparison
Stumbled across this cool visualisation project by Petra Isenberg at Calgary University. Collaborative tree comparison uses a tabletop system to enable two (or more) people to interact when comparing (in this case) phylogenies. I want one!
The system is described in "Interactive Tree Comparison for Co-located Collaborative Information Visualization" (doi:10.1109/TVCG.2007.70568), a PDF of which is available from her web site (which also has a great video entitled "how co-located collaborative data analysis should not take place").
The system is described in "Interactive Tree Comparison for Co-located Collaborative Information Visualization" (doi:10.1109/TVCG.2007.70568), a PDF of which is available from her web site (which also has a great video entitled "how co-located collaborative data analysis should not take place").
Wednesday, September 30, 2009
Visualising the Biodiversity Heritage Library as a Timeline
 One thing about the Encyclopedia of Life which bugs me no end is the awful way it displays the bibliography generated from the Biodiversity Heritage Library (BHL). The image on the right shows the bibliography for the frog Hyla rivularis Taylor, 1952. It's one long, alphabetical list of pages. How can a user make sense of this? It's even more annoying because the BHL is one of the cornerstones of EOL, and one could argue that BHL content is one of the few thing pages EOL offer that distinguishes them from cheap and cheerful mashups such as iSpecies. Can't we do something a little better?
One thing about the Encyclopedia of Life which bugs me no end is the awful way it displays the bibliography generated from the Biodiversity Heritage Library (BHL). The image on the right shows the bibliography for the frog Hyla rivularis Taylor, 1952. It's one long, alphabetical list of pages. How can a user make sense of this? It's even more annoying because the BHL is one of the cornerstones of EOL, and one could argue that BHL content is one of the few thing pages EOL offer that distinguishes them from cheap and cheerful mashups such as iSpecies. Can't we do something a little better?BHL has an API (documented here), so I decided to experiment. As I mentioned in an earlier post (Biodiversity Heritage Library, Google books, and metadata quality), a key piece of metadata about a bibliographic reference is its date. This is especially so for the taxonomic literature, where the earliest reference that contains a name may (depending on how complete BHL scanning is) be the first description of that name. So, it would be nice to order the BHL bibliography by date. Turns out it's possible to get dates from quite a few BHL items, providing one fusses with regular expressions long enough.
So, in principle we could sort BHL content by dates. But, we could go one better and visualise them. As an experiment, I've put together a demo that uses the SIMILE Timeline widget to display the BHL bibliography for a taxon. Here's a screenshot of the bibliography for Hyla rivularis:

You can generate others at http://bioguid.info/bhl/. The demo has been thrown together in haste, but here's what it does:
- Takes a taxon name and tries to find it in uBio. This gives us the NamebankID BHL needs
- Calls the BHL API and retrieves the bibliography for the NamebankID found in step 1
- Parses the BHL result, trying to extract dates (often the dates are ranges, e.g. 1950-1955)
- The previous step generates a JSON document which can be displayed by Timeline
If you click on an item you get a list of pages, clicking on any those takes to you to the page in BHL. Items that have a range of dates are displayed as horizontal lines, items with a well-defined date are shown as points. Note that my code for working out the date of an item will probably fail on some items, and some items don't have any dates at all. Hence, not every item in BHL will appear in the timeline.
It would be nice to embellish the results a little (for example, group pages into articles, refine the dates, etc.) but I think this goes a little way to demonstrating what can be done. We could also add articles obtained from other sources (e.g., Google Scholar, PubMed) to the same display, providing an overview of published knowledge about a taxon.
Labels:
BHL,
bibliographies,
bioguid,
EOL,
JSON,
timeline,
visualisation
Monday, September 28, 2009
Google Scholar metadata quality and Mendeley hype
Hot on the heels of Geoffrey Nunberg's essay about the train wreck that is Google books metadata (see my earlier post) comes Google Scholar’s Ghost Authors, Lost Authors, and Other Problems by Péter Jacsó. It's a fairly scathing look at some of the problems with the quality of Google Scholar's metadata.
Now, Google Scholar isn't perfect, but it's come to play a key role in a variety of bibliographic tools, such as Mendeley, and Papers. These tools do a delicate dance with Google Scholar who, strictly speaking, don't want anybody scraping their content. There's no API, so Mendeley, Papers (and my own iSpecies) have to keep up with the HTML tweaks that Google introduces, pretend to be web browsers, fuss with cookies, and try to keep the rate of queries below the level at which the Google monster stirs and slaps them down.
Jacsó's critique also misses the main point. Why do we have free (albeit closed) tools like Google Scholar in the first place? It's largely because scientists have ceeded the field of citation analysis to commercial companies, such as Elsevier and Thompson Reuters. To echo Martin Kalfatovic's comment:
For me, this is the one thing the ridiculously over-hyped Mendeley could do that would merit the degree of media attention it is getting -- be the basis of an open citation database. It would need massive improvement to its metadata extraction algorithms, which currently suck (Google Scholar's, for all Jacsó's complaints, are much better), but it would generate something of lasting value.
Now, Google Scholar isn't perfect, but it's come to play a key role in a variety of bibliographic tools, such as Mendeley, and Papers. These tools do a delicate dance with Google Scholar who, strictly speaking, don't want anybody scraping their content. There's no API, so Mendeley, Papers (and my own iSpecies) have to keep up with the HTML tweaks that Google introduces, pretend to be web browsers, fuss with cookies, and try to keep the rate of queries below the level at which the Google monster stirs and slaps them down.
Jacsó's critique also misses the main point. Why do we have free (albeit closed) tools like Google Scholar in the first place? It's largely because scientists have ceeded the field of citation analysis to commercial companies, such as Elsevier and Thompson Reuters. To echo Martin Kalfatovic's comment:
Over the years, we've (librarians and the user community) have allowed an important class of metadata - specifically the article level metadata - migrate to for profit entities.Some visionaries, such as Robert Cameron in his A Universal Citation Database as a Catalyst for Reform in Scholarly Communication, argued for free, open citation databases, but this came to nought.
For me, this is the one thing the ridiculously over-hyped Mendeley could do that would merit the degree of media attention it is getting -- be the basis of an open citation database. It would need massive improvement to its metadata extraction algorithms, which currently suck (Google Scholar's, for all Jacsó's complaints, are much better), but it would generate something of lasting value.
Saturday, September 19, 2009
Biodiversity Heritage Library, Google books, and metadata quality
I've been playing recently with the Biodiversity Heritage Library (BHL), and am starting to get a sense for the complexities (and limitations) of the metadata BHL stores about publications. The more I look at BHL the more I think the resource is (a) wonderfully useful and (b) hampered by some dodgy metadata.
The BHL data model has three kinds of entities, "Titles", "Items", and "Pages". Pages are individual pages in an item, where an item which corresponds to a physical object that has been scanned (such as a book or a bound volume of a journal). A title may comprise a single item, such as book, or many items, such as volumes of a journal. Most of the metadata BHL has relates to physical items (books and bound volume issues), as opposed to article-level metadata, which is basically absent (see But where are the articles?).

This model reflects the sources of the BHL metadata (library catalogues) and the mode of operation (bulk scanning of bound volumes). But it can make working out dates of somewhat challenging.
To give an example, I did a search on the frog name Hyla rivularis Taylor, 1952 (NameBankID 27357), currently known as Isthmohyla rivularis. I wanted to find the original description of this frog. A BHL search returns 34 pages containing the name Hyla rivularis, distributed over 5 titles (a title in BHL may be a book, or a journal). Given that the name was published in 1952, it would be nice if I could sort these results by date, and then look at items from 1952. Unfortunately I can't. BHL has limited information on dates, especially at the level I would need to find a document published in 1952.
For the five titles returned in the search, I have dates for four of them, albeit two are ranges (University of Kansas publications, Museum of Natural History, 1946-1971, and The University of Kansas science bulletin, 1902-1996). At the level of individual items, only item 25858 (University of Kansas publications, Museum of Natural History) has dates (1961-1966). If I look at the VolumeInfo field for an item (you can get this from the database dump, or using the JSON web service) I sometimes get strings like this "v.35:pt.1 (1952)". This item (25857) is the one I'm after, but the date is buried in the VolumeInfo string. So, the information I need is there, but it's going to need some parsing.

Another issue is that of duplicates. Searching for publications on Rana grahamii, I found items 41040 and 45847. Although one item is treated as a book, and the other as a volume of the journal Records of the Indian Museum, these are the same thing. Having duplicates is a complication, but it might also be useful for quality control and testing (for example, do taxon name extraction algorithms return the same names from OCR text from both copies?). Nor is having duplicate copies and/or identifiers unique to BHL. The Records of the Indian Museum has a series-level identifier (ISSN 0537-0744), and this article ("A monograph of the South Asian, Papuan, Melanesian and Australian frogs of the genus Rana") also as the ISBN 8121104327.
There are parallels with Google books scanning project, which has been the subject of criticism on several fronts, including the quality of the metadata they have for each book. Geoff Nunberg has an entertaining post entitled Google Books: A Metadata Train Wreck which lists many examples of errors. This blog post also contains a detailed response from Jon Orwant of Google books. In essence, Google books is riddled with metadata errors (such as books on the Internet with publication dates predating the birth of their authors), but most of these errors have come from library catalogues (not unexpected given the scale of the task), not Google.
What could BHL do about its metadata? One thing is crowdsourcing. BHL does a little of this already, for example capturing user-provided metadata when PDFs are created, but I wonder if we could do more. For example, imagine dumping metadata for all 39,000 items into a semantic wiki and inviting people to edit and annotate the metadata. This could be extended to adding article boundaries (i.e., identifying which page corresponds to the start of an article). There is also considerable scope for trying to find article boundaries using existing metadata from bibliographies assembled by individual scientists.
But we should watch closely what Google does with its book project. Eric Hellman has argued that, far from creating the metadata mess, Google is ideally positioned to sort it out. He writes:
The BHL data model has three kinds of entities, "Titles", "Items", and "Pages". Pages are individual pages in an item, where an item which corresponds to a physical object that has been scanned (such as a book or a bound volume of a journal). A title may comprise a single item, such as book, or many items, such as volumes of a journal. Most of the metadata BHL has relates to physical items (books and bound volume issues), as opposed to article-level metadata, which is basically absent (see But where are the articles?).

This model reflects the sources of the BHL metadata (library catalogues) and the mode of operation (bulk scanning of bound volumes). But it can make working out dates of somewhat challenging.
To give an example, I did a search on the frog name Hyla rivularis Taylor, 1952 (NameBankID 27357), currently known as Isthmohyla rivularis. I wanted to find the original description of this frog. A BHL search returns 34 pages containing the name Hyla rivularis, distributed over 5 titles (a title in BHL may be a book, or a journal). Given that the name was published in 1952, it would be nice if I could sort these results by date, and then look at items from 1952. Unfortunately I can't. BHL has limited information on dates, especially at the level I would need to find a document published in 1952.
For the five titles returned in the search, I have dates for four of them, albeit two are ranges (University of Kansas publications, Museum of Natural History, 1946-1971, and The University of Kansas science bulletin, 1902-1996). At the level of individual items, only item 25858 (University of Kansas publications, Museum of Natural History) has dates (1961-1966). If I look at the VolumeInfo field for an item (you can get this from the database dump, or using the JSON web service) I sometimes get strings like this "v.35:pt.1 (1952)". This item (25857) is the one I'm after, but the date is buried in the VolumeInfo string. So, the information I need is there, but it's going to need some parsing.

Another issue is that of duplicates. Searching for publications on Rana grahamii, I found items 41040 and 45847. Although one item is treated as a book, and the other as a volume of the journal Records of the Indian Museum, these are the same thing. Having duplicates is a complication, but it might also be useful for quality control and testing (for example, do taxon name extraction algorithms return the same names from OCR text from both copies?). Nor is having duplicate copies and/or identifiers unique to BHL. The Records of the Indian Museum has a series-level identifier (ISSN 0537-0744), and this article ("A monograph of the South Asian, Papuan, Melanesian and Australian frogs of the genus Rana") also as the ISBN 8121104327.
There are parallels with Google books scanning project, which has been the subject of criticism on several fronts, including the quality of the metadata they have for each book. Geoff Nunberg has an entertaining post entitled Google Books: A Metadata Train Wreck which lists many examples of errors. This blog post also contains a detailed response from Jon Orwant of Google books. In essence, Google books is riddled with metadata errors (such as books on the Internet with publication dates predating the birth of their authors), but most of these errors have come from library catalogues (not unexpected given the scale of the task), not Google.
What could BHL do about its metadata? One thing is crowdsourcing. BHL does a little of this already, for example capturing user-provided metadata when PDFs are created, but I wonder if we could do more. For example, imagine dumping metadata for all 39,000 items into a semantic wiki and inviting people to edit and annotate the metadata. This could be extended to adding article boundaries (i.e., identifying which page corresponds to the start of an article). There is also considerable scope for trying to find article boundaries using existing metadata from bibliographies assembled by individual scientists.
But we should watch closely what Google does with its book project. Eric Hellman has argued that, far from creating the metadata mess, Google is ideally positioned to sort it out. He writes:
What if Google, with pseudo-monopoly funding and the smartest engineers anywhere, manages to figure out new ways to separate the bird shit from the valuable metadata in thousands of metadata feeds, thereby revolutionizing the library world without even intending to do so?
Thursday, September 17, 2009
Towards a wiki of phylogenies
At the start of this week I took part in a biodiversity informatics workshop at the Naturhistoriska riksmuseets, organised by Kevin Holston. It was a fun experience, and Kevin was a great host, going out of his way to make sure myself and other contributors were looked after. I gave my usual pitch along the lines of "if you're not online you don't exist", and talked about iSpecies, identifiers, and wikis.
I also ran a short, not terribly successful exercise using iTaxon to demo what semantic wikis can do. As is often the case with something that hasn't been polished yet, the students found the mechanics of doing things less than intuitive. I need to do a lot of work making data input easier (to date I've focussed on automated adding of data, and forms to edit existing data). Adding data is easy if you know how, but the user needs to know more than they really should have to.
The exercise was to take some frog taxa from the Frost et al. amphibian tree (doi:10.1206/0003-0090(2006)297[0001:TATOL]2.0.CO;2) and link them to GenBank sequences and museum specimens. The hope was that by making these links new information would emerge. You could think of it as an editable version of this. With a bit of post-exercise tidying, we got someway there. The wiki page for the Frost et al.
paper now shows a list of sequences from that paper (not all, I hasten to add), and a map for those sequences that the students added to the wiki:
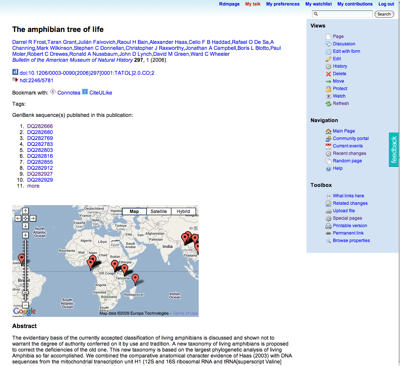
Although much remains to be done, I can't help thinking that this approach would work well for a database like TreeBASE, where one really needs to add a lot of annotation to make it useful (for example, mapping OTUs to taxon names, linking data to sequences and specimens). So, one of the things I'm going to look at is dumping a copy of TreeBASE (complete with trees) into the wiki and seeing what can be done with it. Oh, and I need to make it much, much easier for people to add data.
I also ran a short, not terribly successful exercise using iTaxon to demo what semantic wikis can do. As is often the case with something that hasn't been polished yet, the students found the mechanics of doing things less than intuitive. I need to do a lot of work making data input easier (to date I've focussed on automated adding of data, and forms to edit existing data). Adding data is easy if you know how, but the user needs to know more than they really should have to.
The exercise was to take some frog taxa from the Frost et al. amphibian tree (doi:10.1206/0003-0090(2006)297[0001:TATOL]2.0.CO;2) and link them to GenBank sequences and museum specimens. The hope was that by making these links new information would emerge. You could think of it as an editable version of this. With a bit of post-exercise tidying, we got someway there. The wiki page for the Frost et al.
paper now shows a list of sequences from that paper (not all, I hasten to add), and a map for those sequences that the students added to the wiki:

Although much remains to be done, I can't help thinking that this approach would work well for a database like TreeBASE, where one really needs to add a lot of annotation to make it useful (for example, mapping OTUs to taxon names, linking data to sequences and specimens). So, one of the things I'm going to look at is dumping a copy of TreeBASE (complete with trees) into the wiki and seeing what can be done with it. Oh, and I need to make it much, much easier for people to add data.
Labels:
Mediawiki,
Semantic Web,
TreeBASE,
wiki,
workshop
When taxonomists wage war in Wikipedia
Stumbled across Alex Wild's post Pyramica vs Strumigenys: why does it matter?, which takes as it's starting point a minor edit war on the Wikipedia page for Pyramica .
Alex gives the background to the argument about whether Pyramica is a synonym of Strumigenys, and investigates the issue using the surprisingly small about of data available in GenBank. The tree he found (shown below) suggests this issue will require some work to resolve:

For fun I constructed a history flow diagram for the edits to the Pyramica page in Wikipedia:

The diagram shows the two occasions when the page has been striped of content (and subsequently restored) as contributors dispute whether Pyramica is a synonym of Strumigenys. It would be useful to have one or more metrics of how controversial a page (and/or a contributor) was, to both identify controversial pages, and to see how controversial taxonomic pages were compared to other Wikipedia topics. The paper On Ranking Controversies in Wikipedia: Models and Evaluation by Ba-Quy Vuong et al. (doi:10.1145/1341531.1341556) would be a good place to start (a video of the presentation of this paper is available here).
Alex gives the background to the argument about whether Pyramica is a synonym of Strumigenys, and investigates the issue using the surprisingly small about of data available in GenBank. The tree he found (shown below) suggests this issue will require some work to resolve:

For fun I constructed a history flow diagram for the edits to the Pyramica page in Wikipedia:

The diagram shows the two occasions when the page has been striped of content (and subsequently restored) as contributors dispute whether Pyramica is a synonym of Strumigenys. It would be useful to have one or more metrics of how controversial a page (and/or a contributor) was, to both identify controversial pages, and to see how controversial taxonomic pages were compared to other Wikipedia topics. The paper On Ranking Controversies in Wikipedia: Models and Evaluation by Ba-Quy Vuong et al. (doi:10.1145/1341531.1341556) would be a good place to start (a video of the presentation of this paper is available here).
Labels:
ants,
history flow,
Pyramica,
Strumigenys,
Wikipedia
Friday, September 11, 2009
Gene Wiki and Google
Andrew Su has posted an analysis of Gene Wiki, a project to provide Wikipedia pages on every human gene:
Here's the take home message: in terms of online gene annotation resources, Gene Cards is the most common top-ranked resource, followed closely by the Gene Wiki / Wikipedia, with NCBI in a very distant third (note the log scale).This result is interesting in that an existing resource (Gene Cards) beats Wikipedia, but only just. There are various ways we could interpret this, but from the point of view of biodiversity resources I suspect it emphasises that if there is a good, existing resource that has a lot of traction (i.e., Gene Cards) it will do well in Google Searches. If there is no single dominant resource (as is the case for biodiversity), then it leaves the field open to be dominated by Wikipedia.
Friday, September 04, 2009
Visualising edit history of a Wikipedia page
Quick post (really should be doing something else). Reading Jeff Atwood's post Mixing Oil and Water: Authorship in a Wiki World lead me to IBM's wonderful history flow tool to visualise the edit history of a Wikipedia page.
There's a nice paper describing history flow (doi:10.1145/985692.985765, free PDF here). Inspired by this I decided to try and implement history flow in PHP and SVG. Here's a preliminary result:
This is the edit history for the Afrotheria page. Click on the image above (or here to see the SVG image -- you need a decent web browser for this, IE uses will need a SVG plugin).
The SVG image is clickable. The columns represent revisions, click on those to go to that revision. The columns are evenly spaced (i.e., the gaps don't correspond to time). The bands between revisions trace individual blocks of text (in this case lines in the Wikipedia page source). If you click on a band you get taken to that Wikipedia user's page.
This is all done in a rush, but it gives an idea of what can be done. The history flow carries all sorts of information about how an article has developed over time, major changes (such as the introduction of Taxoboxes), and makes the content of a page traceable, in the sense that you can see who contributed what to a page.
Imagine a scenario where three people will make contributions to a Wiki page at different points in time. Each person edits the page and then saves their changes to what becomes the latest version of that page.
History Flow connects text that has been kept the same between consecutive versions. Pieces of text that do not have correspondence in the next (or previous) version are not connected and the user sees a resulting "gap" in the visualization; this happens for deletions and insertions. (animated GIF from Jeff Atwood's post).
There's a nice paper describing history flow (doi:10.1145/985692.985765, free PDF here). Inspired by this I decided to try and implement history flow in PHP and SVG. Here's a preliminary result:
This is the edit history for the Afrotheria page. Click on the image above (or here to see the SVG image -- you need a decent web browser for this, IE uses will need a SVG plugin).
The SVG image is clickable. The columns represent revisions, click on those to go to that revision. The columns are evenly spaced (i.e., the gaps don't correspond to time). The bands between revisions trace individual blocks of text (in this case lines in the Wikipedia page source). If you click on a band you get taken to that Wikipedia user's page.
This is all done in a rush, but it gives an idea of what can be done. The history flow carries all sorts of information about how an article has developed over time, major changes (such as the introduction of Taxoboxes), and makes the content of a page traceable, in the sense that you can see who contributed what to a page.
Labels:
history flow,
SVG,
visualisation,
Wikipedia
Thursday, September 03, 2009
Google and Wikipedia revisited
Given that one response to my post on Fungi in Wikipedia was to say that fungi are also charismatic, so maybe I should try [insert unsexy taxon name here]. So, I've now looked at all the species I extracted from Wikipedia (nearly 72,000), ran the Google searches, and here are the results:
The table lists the top twenty sites, based on the number of times each site occupies the number one place in the Google search results. Surprise, surprise, Wikipedia wins hands down.
What is interesting is that the other top-ranking sites tend to be taxon-specific, such as FishBase, Amphibia Web, and USDA Plants. To me this suggests that the argument that Wikipedia's dominance of the search results is because it focusses on charismatic taxa doesn't hold. In fact, the truly charismatic taxa are likely to have their own, richly informative webs sites that will often beat Wikipedia in the search rankings. If your taxon is not charismatic, then it's a different story. This suggests one of two strategies for making taxon web sites that people will find. Either go for the niche market, and make a rich site for a set of taxa that you (and ideally some others) like, or add content to Wikipedia. Sites that span across all taxa will always come up against Wikipedia's dominance in the search rankings. So, it's a choice of being a specialist, or trying to compete with an über-generalist.
| Site | How many times is it the top hit? |
|---|---|
| en.wikipedia.org | 42515 |
| www.birdlife.org | 2125 |
| commons.wikimedia.org | 1522 |
| plants.usda.gov | 1496 |
| species.wikimedia.org | 1487 |
| animaldiversity.ummz.umich.edu | 1419 |
| amphibiaweb.org | 851 |
| www.calflora.org | 770 |
| www.fishbase.org | 727 |
| ibc.lynxeds.com | 699 |
| davesgarden.com | 659 |
| www.arkive.org | 510 |
| ukmoths.org.uk | 414 |
| zipcodezoo.com | 368 |
| www.itis.gov | 304 |
| calphotos.berkeley.edu | 294 |
| www.floridata.com | 234 |
| www.planetcatfish.com | 234 |
| www.eol.org | 226 |
| www.arthurgrosset.com | 213 |
The table lists the top twenty sites, based on the number of times each site occupies the number one place in the Google search results. Surprise, surprise, Wikipedia wins hands down.
What is interesting is that the other top-ranking sites tend to be taxon-specific, such as FishBase, Amphibia Web, and USDA Plants. To me this suggests that the argument that Wikipedia's dominance of the search results is because it focusses on charismatic taxa doesn't hold. In fact, the truly charismatic taxa are likely to have their own, richly informative webs sites that will often beat Wikipedia in the search rankings. If your taxon is not charismatic, then it's a different story. This suggests one of two strategies for making taxon web sites that people will find. Either go for the niche market, and make a rich site for a set of taxa that you (and ideally some others) like, or add content to Wikipedia. Sites that span across all taxa will always come up against Wikipedia's dominance in the search rankings. So, it's a choice of being a specialist, or trying to compete with an über-generalist.
Wednesday, September 02, 2009
Fungi in Wikipedia
One response to the analysis I did of the Google rank of mammal pages in Wikipedia is to suggest that Wikipedia does well for mammals because these are charismatic. It's been suggested that for other groups of taxa Wikipedia might not be so prominent in the search results.
As a quick test I extracted the 1552 fungal species I could find in Wikipedia and repeated the analysis. If anything, the results are more dramatic:

Once again, Wikipedia dominates the search rankings. Over 75% of the pages are the top hit in Google. More specialist fungal sites, such as CAB Abstracts Plus and the American Phytopathological Society's online database do pretty well. EOL and the nomenclatural database Index Fungorum barely make an appearance.
If fungi are less "charismatic" than mammals, the implication is that the less charismatic the taxon, the better Wikipedia does (perhaps there is less competition from other sites). Of course, Wikipedia is severely underpopulated with fungal pages, so one could argue that for fungi not in Wikipedia, sites like EOL may do better (relative to other sites), but that would need to be tested. I suspect that sites that provide more broadly useful information (such as APSnet) will continue to dominate the search rankings, followed by scientific articles (for the fungi in Wikipedia the publishers Springer, Wiley, and Elsevier all appear in the top of sites that appear in the Google rankings).
As a quick test I extracted the 1552 fungal species I could find in Wikipedia and repeated the analysis. If anything, the results are more dramatic:

Once again, Wikipedia dominates the search rankings. Over 75% of the pages are the top hit in Google. More specialist fungal sites, such as CAB Abstracts Plus and the American Phytopathological Society's online database do pretty well. EOL and the nomenclatural database Index Fungorum barely make an appearance.
If fungi are less "charismatic" than mammals, the implication is that the less charismatic the taxon, the better Wikipedia does (perhaps there is less competition from other sites). Of course, Wikipedia is severely underpopulated with fungal pages, so one could argue that for fungi not in Wikipedia, sites like EOL may do better (relative to other sites), but that would need to be tested. I suspect that sites that provide more broadly useful information (such as APSnet) will continue to dominate the search rankings, followed by scientific articles (for the fungi in Wikipedia the publishers Springer, Wiley, and Elsevier all appear in the top of sites that appear in the Google rankings).
Tuesday, September 01, 2009
Wikipedia mammals and the power law
Playing a bit more with the Wikipedia mammal data, there are some interesting patterns to note. The first is that rank the mammal pages by size (here defined as the number of characters in the source for the page) and plot size against rank then we get a graph that looks very much like a power law:

There are a few large pages on mammals (these are on the left), and lots of small pages (the long tail on the right). If we do a log-log plot we get this:

The straight line is characteristic of a power law. The dip at the far right reflects the fact that Wikipedia pages have a minimum size (for example, they must include a Taxobox). Now, this is a bit crude (I should probably look at "Power-law distributions in empirical data" arXiv:0706.1062v2 before getting too carried away), but power laws are characteristic of the link structure of the web (a few big sites with huge numbers of links, huge numbers of sites with few links), and indeed of at least parts of Wikipedia, such as the Gene Wiki project (see doi:10.1371/journal.pbio.0060175).
In this context, the diagrams are showing that even if mammals are "charismatic megafauna", most of them aren't that charismatic. Wikipedia mammal pages are mostly small. This raises the question of whether the high frequency in which Wikipedia mammal pages appeared in the top of Google searches might be attributed to those large pages on (presumably) charismatic mammals. If this were the case, then we'd expect that small pages wouldn't rank highly in Google searches. So, I plotted page size against Google search rank for the Wikipedia mammal pages:

This is a box plot, where the grey boxes represent 50% of the distribution of page size (the horizontal black line is the median), and extreme values are shown as circles. Note that "0" is the highest rank (i.e., the first hit in Google), and 9 is the lowest.
While, not surprisingly, most large Wikipedia pages do well in Google searches, and rarely are large pages low down the rankings, my sense is that small pages can have any rank, from top (0) to bottom (9). If page size (i.e., which is a crude measure of the effort put into editing a Wikipedia page) is a measure of "charisma" (contributors are more likely to edit pages on animals that lots of people know about), then charisma isn't a great predictor of where you come in Google's search results. It's not about size, it's about being in Wikipedia.

There are a few large pages on mammals (these are on the left), and lots of small pages (the long tail on the right). If we do a log-log plot we get this:

The straight line is characteristic of a power law. The dip at the far right reflects the fact that Wikipedia pages have a minimum size (for example, they must include a Taxobox). Now, this is a bit crude (I should probably look at "Power-law distributions in empirical data" arXiv:0706.1062v2 before getting too carried away), but power laws are characteristic of the link structure of the web (a few big sites with huge numbers of links, huge numbers of sites with few links), and indeed of at least parts of Wikipedia, such as the Gene Wiki project (see doi:10.1371/journal.pbio.0060175).
In this context, the diagrams are showing that even if mammals are "charismatic megafauna", most of them aren't that charismatic. Wikipedia mammal pages are mostly small. This raises the question of whether the high frequency in which Wikipedia mammal pages appeared in the top of Google searches might be attributed to those large pages on (presumably) charismatic mammals. If this were the case, then we'd expect that small pages wouldn't rank highly in Google searches. So, I plotted page size against Google search rank for the Wikipedia mammal pages:

This is a box plot, where the grey boxes represent 50% of the distribution of page size (the horizontal black line is the median), and extreme values are shown as circles. Note that "0" is the highest rank (i.e., the first hit in Google), and 9 is the lowest.
While, not surprisingly, most large Wikipedia pages do well in Google searches, and rarely are large pages low down the rankings, my sense is that small pages can have any rank, from top (0) to bottom (9). If page size (i.e., which is a crude measure of the effort put into editing a Wikipedia page) is a measure of "charisma" (contributors are more likely to edit pages on animals that lots of people know about), then charisma isn't a great predictor of where you come in Google's search results. It's not about size, it's about being in Wikipedia.
Google, Wikipedia, and EOL
One assumption I've been making so far is that when people search for information on an organism using its scientific name, Wikipedia will dominate the search results (see my earlier post for an example of this assumption). I've decided to quantify this by doing a little experiment. I grabbed the Mammal Species of the World taxonomy and extracted the 5416 species names. I then used Google's AJAX search API to look up each name in Google. For each search I took the top 10 hits and recorded for each hit the site URL and the rank in the search results (i.e., 1-10). Below is a table of how many mammal species had a hit in the top 10 Google results (showing just the top 20 most frequent sites).
Wikipedia is the clear winner, with 5266 (97%) of mammals having a Wikipedia page in the top ten Google results. Next comes Wikispecies, then Animal Diversity Web, Wikimedia Commons, ITIS, the Comparative Toxicogenomics Database, BioOne, UniProt (derived from the NCBI taxonomy), and so on. Note that the Encyclopedia of Life comes in 17th.
Things get more interesting if we look at the ranking of search results. The graph below plots the cumulative rank of search results for some of the web sites listed above.

Wikipedia dominates things. For 48% of all mammal species Wikipedia is the first result returned by Google. Just under three quarters of all mammal species are either the first or second top hit in Google. The next best sites are Animal Diversity Web and Wikispecies, which get a small share of first place for some species (19% and 7% respectively). Note that EOL pages manage to make it into the top 10 for only 11% of all mammal species.
What does this all mean? Well, it seems clear that if people are using Google to find information about an organism, then Wikipedia is more likely than anything else to be the first result they see. It is also interesting that for all the energy (and funds) being expended on biodiversity databases (doi:10.1126/science.324_1632), ITIS is the only classical biodiversity database that routinely gets found in these searches (albeit in only a quarter of the searches).
I know I tend to go on a bit about EOL, but if I was running (or funding) EOL, I'd be worried. EOL barely figures in these search results, and is being taken to the cleaners by a volunteer effort (Wikipedia). Furthermore, it seems difficult to envisage what EOL can do to improve things. Sure it can link to (and make use of) content in sites such as Animal Diversity Web, ITIS (and maybe even, gasp, Wikipedia), but that just adds "link love" to those sites. Ironically, perhaps the single thing that would improve EOL's ranking would be if Wikipedia spread some of its link love over EOL, by linking all it's taxon pages to the corresponding EOL page.
But there are bigger issues at stake. Site popularity on the web tends to follow a power law, where a very few web sites grab the vast majority of eye balls. In a old blog post Clay Shirky wrote:
So, creating new and improved biodiversity web sites is likely to have the effect of only increasing the gap between Wikipedia and the rest.
Lastly, as I've mentioned before regarding Wikipedia and citations of taxonomic work, the graph above suggests to me that for anybody wanting to make basic biodiversity information available on the web, and attract readers to basic taxonomic literature, there really is only one game in town.
| Site | Hits |
|---|---|
| en.wikipedia.org | 5266 |
| species.wikimedia.org | 2934 |
| animaldiversity.ummz.umich.edu | 2890 |
| commons.wikimedia.org | 1515 |
| www.itis.gov | 1418 |
| ctd.mdibl.org | 1288 |
| www.bioone.org | 1101 |
| www.uniprot.org | 1086 |
| encyclopedia.farlex.com | 1007 |
| www.thewebsiteofeverything.com | 955 |
| www.answers.com | 864 |
| vertebrates.si.edu | 854 |
| www.interaktv.com | 842 |
| www.arkive.org | 775 |
| linkinghub.elsevier.com | 727 |
| www.springerlink.com | 656 |
| www.eol.org | 618 |
| www.reference.com | 576 |
| doi.wiley.com | 572 |
| noctilio.com | 566 |
Wikipedia is the clear winner, with 5266 (97%) of mammals having a Wikipedia page in the top ten Google results. Next comes Wikispecies, then Animal Diversity Web, Wikimedia Commons, ITIS, the Comparative Toxicogenomics Database, BioOne, UniProt (derived from the NCBI taxonomy), and so on. Note that the Encyclopedia of Life comes in 17th.
Things get more interesting if we look at the ranking of search results. The graph below plots the cumulative rank of search results for some of the web sites listed above.

Wikipedia dominates things. For 48% of all mammal species Wikipedia is the first result returned by Google. Just under three quarters of all mammal species are either the first or second top hit in Google. The next best sites are Animal Diversity Web and Wikispecies, which get a small share of first place for some species (19% and 7% respectively). Note that EOL pages manage to make it into the top 10 for only 11% of all mammal species.
What does this all mean? Well, it seems clear that if people are using Google to find information about an organism, then Wikipedia is more likely than anything else to be the first result they see. It is also interesting that for all the energy (and funds) being expended on biodiversity databases (doi:10.1126/science.324_1632), ITIS is the only classical biodiversity database that routinely gets found in these searches (albeit in only a quarter of the searches).
I know I tend to go on a bit about EOL, but if I was running (or funding) EOL, I'd be worried. EOL barely figures in these search results, and is being taken to the cleaners by a volunteer effort (Wikipedia). Furthermore, it seems difficult to envisage what EOL can do to improve things. Sure it can link to (and make use of) content in sites such as Animal Diversity Web, ITIS (and maybe even, gasp, Wikipedia), but that just adds "link love" to those sites. Ironically, perhaps the single thing that would improve EOL's ranking would be if Wikipedia spread some of its link love over EOL, by linking all it's taxon pages to the corresponding EOL page.
But there are bigger issues at stake. Site popularity on the web tends to follow a power law, where a very few web sites grab the vast majority of eye balls. In a old blog post Clay Shirky wrote:
Now, thanks to a series of breakthroughs in network theory by researchers ... we know that power law distributions tend to arise in social systems where many people express their preferences among many options. We also know that as the number of options rise, the curve becomes more extreme. This is a counter-intuitive finding - most of us would expect a rising number of choices to flatten the curve, but in fact, increasing the size of the system increases the gap between the #1 spot and the median spot.
So, creating new and improved biodiversity web sites is likely to have the effect of only increasing the gap between Wikipedia and the rest.
Lastly, as I've mentioned before regarding Wikipedia and citations of taxonomic work, the graph above suggests to me that for anybody wanting to make basic biodiversity information available on the web, and attract readers to basic taxonomic literature, there really is only one game in town.
Labels:
Clay Shirky,
EOL,
Google,
power law,
search,
Wikipedia,
Wikispecies
Monday, August 31, 2009
Comparing Wikipedia and Mammal Species of the World classifications
Continuing the saga of making sense of the mammal classification in Wikipedia, I've done a quick comparison with the Mammal Species of the World (third edition) classification. MSW is the default taxonomic reference used by WikiProject Mammals. I downloaded the MSW taxonomy as a CSV file (warning, it's big), and wrote a script to pull out the classification as a GML file (my preferred graph format).
Based on some earlier work with Gabriel Valiente, I wrote a simple program that takes two trees and highlights the nodes in common to the two trees. I then input into this program the MSW tree, and the largest component of the graph of Wikipedia mammals. The MSW tree has 13582 nodes, the Wikipedia tree has 6287. Note that Wikipedia has more taxa than these 6287 nodes suggest, but they aren't connected to the largest tree (often due to intermediate nodes in the classification lacking a page in Wikipedia). The two trees have 4935 nodes in common (again, this number will be a little low, there are some weird taxon names due to problems parsing Wikipedia).
MSW versus Wikipedia
Below is a the MSW classification with taxa in Wikipedia shown in red.

[Larger scale view here]
The impression given is that most Wikipedia mammal pages are in MSW, with some notable exceptions, including higher level taxa such as Afrotheria, and extinct taxa such as the Multituberculata. Some extant taxa are missing due to synonymy. For example, Wikipedia gives the scientific name of Anthony's pipistrelle as Pipistrellus anthonyi, whereas MSW has it as Hypsugo anthonyi.
As an aside, Wikipedia pages often get muddled about parentheses around taxonomic author names. The authority is in parentheses if the current genus is not the original genus the species was placed. Hence, Pipistrellus anthonyi (Tate, 1942) should actually be Pipistrellus anthonyi Tate, 1942, as Tate originally described this taxon as a species of Pipistrellus (see hdl:2246/1783). However, the name Hypsugo anthonyi (Tate, 1942) does need parentheses.
Some Wikipedia taxa also postdate the publication of MSW, such as Philander deltae (see doi:10.1644/05-MAMM-A-065R2.1).
Wikipedia versus MSW
When we do the reverse comparison we see something rather different.

[Larger scale view here]
This is the MSW tree, coloured red where the MSW taxon has a page in Wikipedia. There are big gaps, some of which are due to those pages being in another component (in other words, many "missing" taxa do have pages in Wikipedia, they are just not properly linked to the bigger tree). MSW is also rich in subspecies, which tend to lack their own pages in Wikipedia (possibly a good thing in the cases of taxa such as pocket gophers).
It would be nice to make these comparisons automatic, and develop tools so that managing taxonomy in Wikipedia could be made easier.
Saturday, August 29, 2009
Mammal tree from Wikipedia
Following on from my previous post about visualising the mammalian classification in Wikipedia, I've extracted the largest component from the graph for all mammal taxa in Wikipedia, and it is a tree. This wasn't apparent in the previous diagram, where the component appeared as a big ball due to the layout algorithm used.

What this suggests is that Wikipedia contributors are quite capable of generating trees, it's just that not all the bits of the tree are connected (hence all the components in the previous post.
As Cyndy Parr suggested in her comments, it would be useful to compare the Wikipedia-derived tree with other trees, say from Mammal species of the World or ITIS.

What this suggests is that Wikipedia contributors are quite capable of generating trees, it's just that not all the bits of the tree are connected (hence all the components in the previous post.
As Cyndy Parr suggested in her comments, it would be useful to compare the Wikipedia-derived tree with other trees, say from Mammal species of the World or ITIS.
Friday, August 28, 2009
Visualising the Wikipedia classification of mammals
As part of my on-going experiments with Wikipedia as a repository of taxonomic information, I've extracted mammal pages from Wikipedia. There's a lot to be done with these, but the first thing I wanted to ask was whether the Wikipedia pages would form a tree (i.e., had the authors of these pages managed to ensure the pages formed a single, coherent taxonomic classification). The answer, as shown in the graph below, is no.

The graph contains 7750 nodes, each one representing a Wikipedia page with a Taxobox containing the class Mammalia. A node is connected to the node corresponding to its parent in the mammalian classification.
If it formed a single classification there would be just one component. Instead, it contains 841 distinct components, many of which you can see at the bottom. If you want to explore the graph, I've made an image map here using the wonderful graph editor yEd. You'll need to move the browser's scroll bars to see the graph. If you click on the node you'll be taken to the corresponding Wikipedia page.
Note: The graph has been laid out using yEd's organic layout command, so it won't look tree-like. The diagram is intended to testing for connectedness only.
Some of these components may be due to errors in my parser, but many are due to inconsistencies in Wikipedia. Typical problems are Taxoboxes containing taxa for which there is no page in Wikipedia (these are visible as redlinks), or monotypic taxa where the pages for the genus and species are the same).
Of course, the joy of Wikipedia is that these problems can be easily fixed, but the trick is discovering the problems in the first place. There is a distinct lack of tools to enable Wikipedia editors to view the entire classification of interest and identify areas that need fixing (something Roger Hyam alluded to in his comment on an earlier posting). It would, of course, be great to be able to edit the graph shown above and have those changes automatically transmitted to Wikipedia.

The graph contains 7750 nodes, each one representing a Wikipedia page with a Taxobox containing the class Mammalia. A node is connected to the node corresponding to its parent in the mammalian classification.
If it formed a single classification there would be just one component. Instead, it contains 841 distinct components, many of which you can see at the bottom. If you want to explore the graph, I've made an image map here using the wonderful graph editor yEd. You'll need to move the browser's scroll bars to see the graph. If you click on the node you'll be taken to the corresponding Wikipedia page.
Note: The graph has been laid out using yEd's organic layout command, so it won't look tree-like. The diagram is intended to testing for connectedness only.
Some of these components may be due to errors in my parser, but many are due to inconsistencies in Wikipedia. Typical problems are Taxoboxes containing taxa for which there is no page in Wikipedia (these are visible as redlinks), or monotypic taxa where the pages for the genus and species are the same).
Of course, the joy of Wikipedia is that these problems can be easily fixed, but the trick is discovering the problems in the first place. There is a distinct lack of tools to enable Wikipedia editors to view the entire classification of interest and identify areas that need fixing (something Roger Hyam alluded to in his comment on an earlier posting). It would, of course, be great to be able to edit the graph shown above and have those changes automatically transmitted to Wikipedia.
Friday, August 21, 2009
Scientific citations in Wikipedia

While thinking about measuring the quality of Wikipedia articles by counting the number of times they cite external literature, and conversely measuring the impact of papers by how many times they're cited in Wikipedia, I discovered, as usual, that somebody has already done it. I came across this nice paper by Finn Årup Nielsen (arXiv:0705.2106v1) (originally published in First Monday as a HTML document, I've embedded the PDF from arXiv below).
Nielsen retrieved 30,368 citations from Wikipedia, and summarised how many times each journal is cited within Wikipedia. He then compared this with a measure of citations within the scientific literature by multiplying the journal's impact factor by the total number of citations. In general there's a pretty good correlation.

What is striking to me is that
When individual journals are examined Wikipedia citations to astronomy journals stand out compared to the overall trend (Figure 2). Also Australian botany journals received a considerable number of citations, e.g., Nuytsia (101 [citations]), in part due to concerted effort for the genus Banksia, where several Wikipedia articles for Banksia species have reached "featured article" status.
In the diagram, note also that Australian Systematic Botany (ISSN 1030-1887), which has a impact factor of 1.351, is punching well above its weight in Wikipedia. What I want to find out is whether this is true for other taxonomic journals. Nielsen's study was based on a Wikipedia dump from 2 April 2007, and a lot has been added since then (and the journal Zootaxa has become a major publisher of new taxonomic names).
But what I'm also wondering is whether this is not a great opportunity for the taxonomic community. By responding to {{citation needed}}, we can improve the quality of Wikipedia, and increase the visibility of their work. Given that many Wikipedia taxon pages are in the top 10 Google hits {{citation needed}}, our work is but one click away from the Google results page. Instead of endlessly moaning about the low impact factor of taxonomic journals, we can actively do something that increases the quality and visibility of taxonomic information, and by extension, taxonomy itself.
Scientific citations in Wikipedia
Tuesday, August 18, 2009
To wiki or not to wiki?
What follows are some random thoughts as I try and sort out what things I want to focus on in the coming days/weeks. If you don't want to see some wallowing and general procrastination, look away now.
I see four main strands in what I've been up to in the last year or so:
Services
Not glamourous, but necessary. This is basically bioGUID (see also hdl:10101/npre.2009.3079.1). bioGUID provides OpenURL services for resolving articles (it has nearly 84,000 articles in it's cache), looking up journal names, resolving LSIDs, and RSS feeds.
Mashups
iSpecies is my now aging tool for mashing up data from diverse sources, such as Wikipedia, NCBI, GBIF, Yahoo, and Google Scholar. I tweak it every so often (mainly to deal with Google Scholar forever mucking around with their HTML). The big limitation of iSpecies is that it doesn't make it's results reusable (i.e., you can't write a script to call iSpecies and return data). However, it's still the place I go to to quickly find out about a taxon.
The other mashups I've been playing with focus on taking standardised RSS feeds (provided by bioGUID, see above) and mashing them up, sometimes with a nice front end (e.g., my e-Biosphere 09 challenge entry).
Wiki
I've invested a huge amount of effort in learning how wikis (especially Mediawiki and its semantic extensions) work, documented in earlier posts. I created a wiki of taxonomic names as a sandbox to explore some of these ideas.
I've come to the conclusion that for basic taxonomic and biological information, the only sensible strategy for our community is to use (and contribute to) Wikipedia. I'm struggling to see any justification for continuing with a proliferation of taxonomic databases. After e-Biosphere 09 the game's up, people have started to notice that we've an excess of databases (see Claire Thomas in Science, "Biodiversity Databases Spread, Prompting Unification Call", doi:10.1126/science.324_1632).
Phyloinformatics
In truth I've not been doing much on this, apart from releasing tvwidget (code available from Google Code), and playing with a mapping of TreeBASE studies to bibliographic identifiers (available as a featured download from here). I've played with tvwidget in Mediawiki, and it seems to work quite well.
Where now?
So, where now? Here are some thoughts:
I see four main strands in what I've been up to in the last year or so:
- services
- mashups
- wikis
- phyloinformatics
Services
Not glamourous, but necessary. This is basically bioGUID (see also hdl:10101/npre.2009.3079.1). bioGUID provides OpenURL services for resolving articles (it has nearly 84,000 articles in it's cache), looking up journal names, resolving LSIDs, and RSS feeds.
Mashups
iSpecies is my now aging tool for mashing up data from diverse sources, such as Wikipedia, NCBI, GBIF, Yahoo, and Google Scholar. I tweak it every so often (mainly to deal with Google Scholar forever mucking around with their HTML). The big limitation of iSpecies is that it doesn't make it's results reusable (i.e., you can't write a script to call iSpecies and return data). However, it's still the place I go to to quickly find out about a taxon.
The other mashups I've been playing with focus on taking standardised RSS feeds (provided by bioGUID, see above) and mashing them up, sometimes with a nice front end (e.g., my e-Biosphere 09 challenge entry).
Wiki
I've invested a huge amount of effort in learning how wikis (especially Mediawiki and its semantic extensions) work, documented in earlier posts. I created a wiki of taxonomic names as a sandbox to explore some of these ideas.
I've come to the conclusion that for basic taxonomic and biological information, the only sensible strategy for our community is to use (and contribute to) Wikipedia. I'm struggling to see any justification for continuing with a proliferation of taxonomic databases. After e-Biosphere 09 the game's up, people have started to notice that we've an excess of databases (see Claire Thomas in Science, "Biodiversity Databases Spread, Prompting Unification Call", doi:10.1126/science.324_1632).
Phyloinformatics
In truth I've not been doing much on this, apart from releasing tvwidget (code available from Google Code), and playing with a mapping of TreeBASE studies to bibliographic identifiers (available as a featured download from here). I've played with tvwidget in Mediawiki, and it seems to work quite well.
Where now?
So, where now? Here are some thoughts:
- I will continue to hack bioGUID (it's now consuming RSS feeds from journals, as well as Zotero). Everything I do pretty much depends on the services bioGUID provides
- iSpecies really needs a big overhaul to serve data in a form that can be built upon. But this requires decisions on what that format should be, so this isn't likely to happen soon. But I think the future of mashup work is to use RDF and triple stores (providing that some degree of editing is possible). I think a tool linking together different data sources (along the lines of my ill-fated Elsevier Challenge entry) has enormous potential.
- I'm exploring Wikipedia and Wikispecies. I'm tempted to do a quantitative analysis of Wikipedia's classification. I think there needs to be some serious analysis of Wikipedia if people are going to use it as a major taxonomic resource.
- If I focus on Wikipedia (i.e., using an existing wiki rather than try to create my own), then that leaves me wondering what all the playing with iTaxon was for. Well, actually I think the original goal of this blog (way back in December 2005) is ideally suited to a wiki. Pretty much all the elements are in place to dump a copy of TreeBASE into a wiki and open up the editing of links to literature and taxonomic names. I think this is going to handily beat my previous efforts (TbMap, doi:10.1186/1471-2105-8-158), especially as errors will be easy to fix.
Subscribe to:
Posts (Atom)