Continuing the theme of the
failings of the GBIF classification I've been playing further with cluster maps to visualise the problem (see
this earlier post for an introduction).
Browsing through bats in GBIF I keep finding the same species appearing more than once, albeit in different genera. As discussed in the
gibbon example, GBIF merges several competing classifications for mammals, and these often don't agree on the "accepted name" for a species. In the absence of a decent database of taxonomic synonyms, GBIF ends up duplicating species, and each duplicate is often associated with different occurence data. If you are trying to get the distribution for a species this can be a disaster.
To get a sense of the scale of the problem I put together a simple tool to create cluster maps. The code is on
github) and there is a live service at
http://iphylo.org/~rpage/cluster-map/. The service takes a simple tab-delimited file that lists sets and their members, computes the overlap between the sets, calls Graphviz to layout a graph in SVG, then draws in the members of each cluster (phew).
The input file looks something like this:
Molossops aequatorianus
Chaerephon aloysiisabaudiae
Tadarida aloysiisabaudiae
Chaerephon ansorgei
Tadarida ansorgei
Molossus ater
Mormopterus petrophilus
Sauromys petrophilus
What can we do with this tool? Well, I created a quick list of all the species of bat in the family Molossidae according to GBIF. The sets are the bat genera, the members are the species (you can see the file
here). I then ran this through the cluster map, and got something like this (this is only part of the cluster map):

(now can you see why I call these "papaya plots"?). Note that there are species names (i.e., specific epithets) in common to more than one genus. Some of these may be perfectly OK (it's not unusual for the same epithet to be used in different species, e.g. "major", etc.). But in many cases these bat species turn out to be
the same species, just in different genera in different classifications. For example, GBIF has both
Cynomops greenhalli and
Molossops greenhalli. These are the same thing. Species in the genus
Mormopterus may also occur in other genera. In some cases the issue is competing classifications, sometimes it is conflict over whether a species is a species or merely a subspecies, and some generic conflicts are because some genera are relegated to subgeneric status in some classifications. In short, it's an unholy mess.
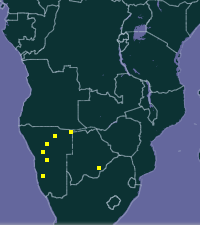
Does this matter? Well, consider
Mormopterus petrophilus and
Sauromys petrophilus, which GBIF both regard as valid species (they're the same thing). Here are the distributions for the two different names in GBIF:
Depending on which name you use you'll get a very different picture of the distribution of this bat.
The next step is to figure out how to fix this. Is there a way we can automate fixing the GBIF classification so that it is not riddled with spurious duplicates like these?