More arm-waving notes on taxonomic databases. I've started to add data to ChecklistBank and this has got me thinking about the issue of data quality. When you add data to ChecklistBank you are asked to give a measure of confidence based on the Catalogue of Life Checklist Confidence system of one - five stars: ★ - ★★★★★. I'm scepetical about the notion of confidence or "trust" when it is reduced to a star system (see also Can you trust EOL?). I could literally pick any number of stars, there's no way to measure what number of stars is appropriate. This feeds into my biggest reservation about the Catalogue of Life, it's almost entirely authority based, not evidence based. That is, rather than give us evidence for why a particular taxon is valid, we are (mostly) just given a list of taxa are asked to accept those as gospel, based on assertions by one or more authorities. I'm not necessarly doubting the knowledge of those making these lists, it's just that I think we need to do better than "these are the accepted taxa because I say so" implict in the Catalogue of Life.
So, is there any way we could objectively measure the quality of a particular taxonomic checklist? Since I have a long standing interest in link the primary taxonomic litertaure to names in databases (since that's where the evidence is), I keep wondering whether measures based on that literture could be developed.
I recently revisited the fascinating (and quite old) literature on rates of synonymy:
Gaston Kevin J. and Mound Laurence A. 1993 Taxonomy, hypothesis testing and the biodiversity crisisProc. R. Soc. Lond. B.251139–142 http://doi.org/10.1098/rspb.1993.0020
Andrew R. Solow, Laurence A. Mound, Kevin J. Gaston, Estimating the Rate of Synonymy, Systematic Biology, Volume 44, Issue 1, March 1995, Pages 93–96, https://doi.org/10.1093/sysbio/44.1.93
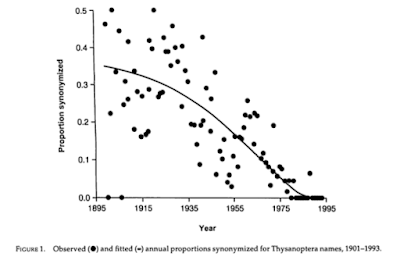
A key point these papers make is that the observed rate of synonymy is quite high (that is, many "new species" end up being merged with already known species), and that because it can take time to discover that a species is a synonym the actual rate may be even higher. In other words, in diagrams like the one reproduced below, the reason the proportion of synonyms declines the nearer we get to the present day (this paper came out in 1995) is not because are are creating fewer synonyms but because we've not yet had time to do the work to uncover the remaining synonyms.
Put another way, these papers are arguing that real work of taxonomy is revision, not species discovery, especially since it's not uncommon for > 50% of species in a taxon to end up being synonymised. Indeed, if a taxoomic group has few synonyms then these authors would argue that's a sign of neglect. More revisionary work would likely uncover additional synonyms. So, what we need is a way to measure the amount of research on a taxonomic group. It occurs to me that we could use the citation graph as a way to tackle this. Lets imagine we have a set of taxa (say a family) and we have all the papers that described new species or undertook revisions (or both). The extensiveness of that work could be measured by the citation graph. For example, build the citation graph for those papers. How many original species decsriptions are not cited? Those species have been potentially neglected. How many large-scale revisions have there been (as measured by the numbers of taxonomic papers those revisions cite)? There are some interesting approaches to quantifying this, such as using hubs and authorities.
I'm aware that taxonomists have not had the happiest relationship with citations:
Pinto ÂP, Mejdalani G, Mounce R, Silveira LF, Marinoni L, Rafael JA. Are publications on zoological taxonomy under attack? R Soc Open Sci. 2021 Feb 10;8(2):201617. doi: 10.1098/rsos.201617. PMID: 33972859; PMCID: PMC8074659.Still, I think there is an intriguing possibility here. For this approach to work, we need to have linked taxonomic names to publications, and have citation data for those publications. This is happening on various platforms. Wikidata, for example, is becoming a repository of the taxonomic literature, some of it with citation links.
Page RDM. 2022. Wikidata and the bibliography of life. PeerJ 10:e13712 https://doi.org/10.7717/peerj.13712Time for some experiments.