Dario Taraborelli has released ReaderMeter, an elegant app built on top of the Mendeley API. You enter an author's name and it summarises that authorship's readership in Mendeley. The app provides some summary statistics (mine are shown below), and if you click on the horizontal bar corresponding to a paper, you can see a visualisation of who is reading your paper, including a nice map.
Dario Taraborelli has released ReaderMeter, an elegant app built on top of the Mendeley API. You enter an author's name and it summarises that authorship's readership in Mendeley. The app provides some summary statistics (mine are shown below), and if you click on the horizontal bar corresponding to a paper, you can see a visualisation of who is reading your paper, including a nice map.
As ever with author names, there are issues of people's name having more than one spelling. In Mendeley I'm known as Roderic D. M. Page, R. D. M. Page, Rod Page, Roderic Page, Roderic D. M. Page, and doubtless some others. Searching ReaderMeter using different spellings of my name gives different results. There are various approaches to tackling this problem, I've touched on one approach earlier.
However, there's a different way to tackle this problem in the context of apps like ReaderMeter, because if you're a Mendeley user you can assert that you are the author of a paper (these papers live in your "My Publications" collection). Using Mendeley's API, an app could retrieve this list of publications (providing the user gave it access), and we could compute readership statistics from the set of articles "known" to be authored (leaving aside the issue of people gaming the system by spuriously claiming authorship). In this way the app relies on the default behaviour of Mendeley users - uploading and self-identifying the articles they've written.
Implementing a feature like this posses two problems. The first is access to a user's data. Mendeley's API supports OAuth, so it could be done in such a way that only the account's user could authorise the app to access this list. The app could store the fact that the user has verified that the list of publications. Think of it as a bit like Amazon's Real Name™ feature.
The other obstacle is Mendeley's API, which returns readership statistics for public documents (i.e., those in the central aggregation). At present, using the API there is no way to link the global id for a Mendeley reference (e.g., ae7dd6a0-6d09-11df-936c-0026b95e484c) with the local id (e.g., 3582682802) that reference has in a user's collection, unless we resort to trying to match articles by searching by identifiers or article titles. If the API exposed these links, apps like ReaderMeter could become even more powerful (and personalised).



 Continuing the series of posts about reading scientific articles on the iPad, here are some quick notes on perhaps the most polished app I've seen,
Continuing the series of posts about reading scientific articles on the iPad, here are some quick notes on perhaps the most polished app I've seen, 


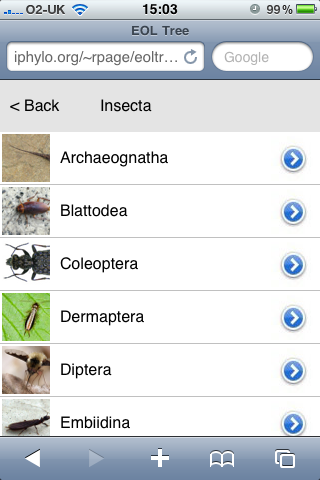 Using the API
Using the API
 Following on from my
Following on from my