There are several instances where I have a collection of references that I want to deduplicate and merge. For example, in Zootaxa has no impact factor I describe a dataset of the literature cited by articles in the journal Zootaxa. This data is available on Figshare (https://doi.org/10.6084/m9.figshare.c.5054372.v4), as is the equivalent dataset for Phytotaxa (https://doi.org/10.6084/m9.figshare.c.5525901.v1). Given that the same articles may be cited many times, these datasets have lots of duplicates. Similarly, articles in Wikispecies often have extensive lists of references cited, and the same reference may appear on multiple pages (for an initial attempt to extract these references see https://doi.org/10.5281/zenodo.5801661 and https://github.com/rdmpage/wikispecies-parser).
There are several reasons I want to merge these references. If I want to build a citation graph for Zootaxa or Phytotaxa I need to merge references that are the same so that I can accurate count citations. I am also interested in harvesting the metadata to help find those articles in the Biodiversity Heritage Library (BHL), and the literature cited section of scientific articles is a potential goldmine of bibliographic metadata, as is Wikispecies.
After various experiments and false starts I've created a repository https://github.com/rdmpage/bib-dedup to host a series of PHP scripts to deduplicate bibliographics data. I've settled on using CSL-JSON as the format for bibliographic data. Because deduplication relies on comparing pairs of references, the standard format for most of the scripts is a JSON array containing a pair of CSL-JSON objects to compare. Below are the steps the code takes.
Generating pairs to compare
The first step is to take a list of references and generate the pairs that will be compared. I started with this approach as I wanted to explore machine learning and wanted a simple format for training data, such as an array of two CSL-JSON objects and an integer flag representing whether the two references were the same of different.
There are various ways to generate CSL-JSON for a reference. I use a tool I wrote (see Citation parsing tool released) that has a simple API where you parse one or more references and it returns that reference as structured data in CSL-JSON.
Attempting to do all possible pairwise comparisons rapidly gets impractical as the number of references increases, so we need some way to restrict the number of comparisons we make. One approach I've explored is the “sorted neighbourhood method” where we sort the references 9for example by their title) then move a sliding window down the list of references, comparing all references within that window. This greatly reduces the number of pairwise comparisons. So the first step is to sort the references, then run a sliding window over them, output all the pairs in each window (ignoring in pairwise comparisons already made in a previous window). Other methods of "blocking" could also be used, such as only including references in a particular year, or a particular journal.
So, the output of this step is a set of JSON arrays, each with a pair of references in CSL-JSON format. Each array is stored on a single line in the same file in line-delimited JSON (JSONL).
Comparing pairs
The next step is to compare each pair of references and decide whether they are a match or not. Initially I explored a machine learning approach used in the following paper:
Wilson DR. 2011. Beyond probabilistic record linkage: Using neural networks and complex features to improve genealogical record linkage. In: The 2011 International Joint Conference on Neural Networks. 9–14. DOI: 10.1109/IJCNN.2011.6033192
Initial experiments using https://github.com/jtet/Perceptron were promising and I want to play with this further, but I deciding to skip this for now and just use simple string comparison. So for each CSL-JSON object I generate a citation string in the same format using CiteProc, then compute the Levenshtein distance between the two strings. By normalising this distance by the length of the two strings being compared I can use an arbitrary threshold to decide if the references are the same or not.
Clustering
For this step we read the JSONL file produced above and record whether the two references are a match or not. Assuming each reference has a unique identifier (needs only be unique within the file) then we can use those identifier to record the clusters each reference belongs to. I do this using a Disjoint-set data structure. For each reference start with a graph where each node represents a reference, and each node has a pointer to a parent node. Initially the reference is its own parent. A simple implementation is to have an array index by reference identifiers and where the value of each cell in the array is the node's parent.
As we discover pairs we update the parents of the nodes to reflect this, such that once all the comparisons are done we have a one or more sets of clusters corresponding to the references that we think are the same. Another way to think of this is that we are getting the components of a graph where each node is a reference and pair of references that match are connected by an edge.
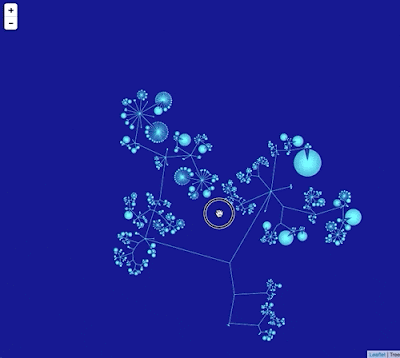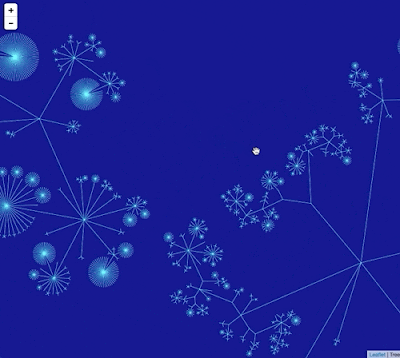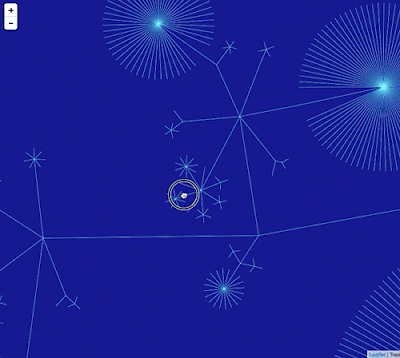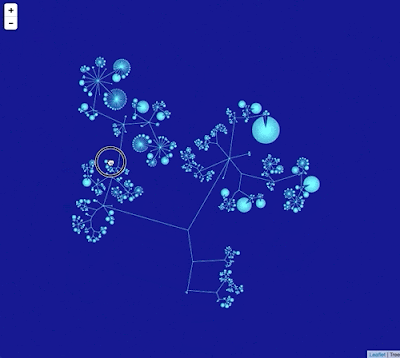
In the code I'm using I write this graph in Trivial Graph Format (TGF) which can be visualised using a tools such as yEd.
Merging
Now that we have a graph representing the sets of references that we think are the same we need to merge them. This is where things get interesting as the references are similar (by definition) but may differ in some details. The paper below describes a simple Bayesian approach for merging records:
Councill IG, Li H, Zhuang Z, Debnath S, Bolelli L, Lee WC, Sivasubramaniam A, Giles CL. 2006. Learning Metadata from the Evidence in an On-line Citation Matching Scheme. In: Proceedings of the 6th ACM/IEEE-CS Joint Conference on Digital Libraries. JCDL ’06. New York, NY, USA: ACM, 276–285. DOI: 10.1145/1141753.1141817.
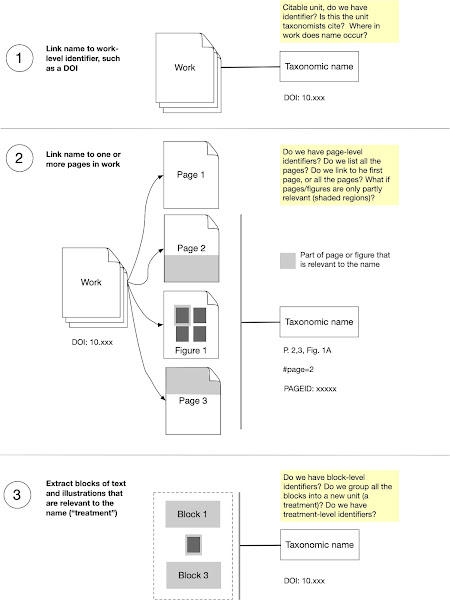
So the next step is to read the graph with the clusters, generate the sets of bibliographic references that correspond to each cluster, then use the method described in Councill et al. to produce a single bibliographic record for that cluster. These records could then be used to, say locate the corresponding article in BHL, or populate Wikidata with missing references.
Obviously there is always the potential for errors, such as trying to merge references that are not the same. As a quick and dirty check I flag as dubious any cluster where the page numbers vary among members of the cluster. More sophisticated checks are possible, especially if I go down the ML route (i.e., I would have evidence for the probability that the same reference can disagree on some aspects of metadata).
Summary
At this stage the code is working well enough for me to play with and explore some example datasets. The focus is on structured bibliographic metadata, but I may simplify things and have a version that handles simple string matching, for example to cluster together different abbreviations of the same journal name.