In an earlier post (
Are names really the key to the big new biology?, I questioned Patterson et al.'s assertion in a recent TREE article (
doi:10.1016/j.tree.2010.09.004) that names are key to the new biology.
In this post I'm going to revisit this idea by doing a quick analysis of how many species in GenBank have "proper" scientific names, and whether the number of named species has changed over time. My definition of "proper" name is a little loose: anything that had two words, second one starting with a lower case letter, was treated as a proper name. hence, a name like
Eptesicus sp. A JLE-2010" is not a proper name, but
Eptesicus andersoni is.
MammalsSince GenBank started, every year has seen some 100-200 mammal species added to the database.

Until around 2003 almost all of these species had proper binomial names, but since then an increasing percentage of species-level taxa haven't been identified to species. In 2010 three-quarters of new tax_ids for mammals weren't identified.
 Invertebrates
InvertebratesFor "invertebrates" 2010 saw an explosive growth in the number of new taxa sequenced, with nearly 71,000 new taxa added to GenBank.

This coincides with a spectacular drop in the number of properly-named taxa, but even before 2010 the proportion of named invertebrate species in GenBank was in decline: in 2009 just over a half of the species added had binomials.
 Bacteria
BacteriaTo put this in perspective, here are the equivalent graphs for bacteria.
Although at the outset most of the bacteria in GenBank had binomial names, pretty quickly the bulk of sequenced bacteria had informal names. In 2010 less than 1% of newly sequenced bacteria had been formerly described.

 Dark taxa
Dark taxaFor bacteria the graphs are hardly surprising. To get a proper name a bacterium must be cultured, and the vast majority of bacteria haven't been (or can't be) cultured. Hence, microbiologists can gloat at the nomenclatural mess plant and animal taxonomists have to deal with only because microbiologists have a tiny number of names to deal with.
For mammals and invertebrates there's clear a decline in the use of proper names.It would be tempting to suggest that this reflects a decline in the number of taxonomists - there might simply not be enough of them in enough groups to be able to identify and/or describe the taxa being sequenced.
However, if we look at the recent peaks of unnamed animal species, we discover that many have names like
Lepidoptera sp. BOLD:AAD7075, indicating that they are DNA Barcodes from the
Barcode of Life Data Systems. Of the 62,365 unnamed invertebrates added last year, 54,546 are BOLD sequences that haven't been assigned to a known species. Of the 277 unnamed mammals, 218 are BOLD taxa. Hence, DNA bnacording is flooding Genbank with taxa that lack proper names (and typically are represented by a single DNA bnacode sequence).
There are various ways to interpret these graphs, but for me the message is clear. The bulk of newly added taxa in GenBank are what we might term "dark taxa", that is, taxa that aren't identified to a known species. This doesn't necessarily mean that they are species new to science, we may already have encountered these species before, they may be sitting in museum collections, and have descriptions already published. We simply don't know. As the output from DNA barcoding grows, the number of dark taxa will only increase, and macroscopic biology starts to look a lot like microbiology.
A post-taxonomic worldIf we look at the graphs for bacteria, we see that taxonomic names are virtually irrelevant, and yet microbiology seems to be doing fine as a discipline. So, perhaps it's time to think about a post-taxonomic world where taxonomic names,
contra Patterson et al., are not that important. We can discover a good deal about organismal biology from GenBank alone (see my post
Visualising the symbiome: hosts, parasites, and the Tree of Life for some examples, as well as Rougerie et al. 2010
doi:10.1111/j.1365-294X.2010.04918.x).
This leaves us with two questions:
- How much biology can we do without taxonomic names?
- If the lack of taxonomic names limits what we can do (and, playing devil's advocate, this is an open question) how can we speed up linking GenBank sequences to names?
I suspect that the answer to (1) is "quite a lot" (especially if we think like microbiologists). Question (2) is ultimately a question about how fast we can link literature, museum collections, sequences, and phylogenies. If progress to date is any indication, we need to rethink how we do this, and in a hurry, because dark taxa are accumulating at an accelerating rate.
How the analyses were doneAlthough the NCBI makes a dump of its taxonomic database available via FTP (at
ftp://ftp.ncbi.nih.gov/pub/taxonomy/), this dump doesn't have dates for when the taxa were added to the database. However, using the
Entrez EUtilities we can get the tax_ids that were published within a given date range. For example, to retrieve all the tax_ids added to the database in December 2010, we set the URL parameters
&mindate=2010/12/01 and
&maxdate=2010-12-31 to form this URL:
http://eutils.ncbi.nlm.nih.gov/entrez/eutils/esearch.fcgi?db=taxonomy&mindate=2010/12/01&maxdate=2010/12/31&retmax=1000000.
I've set
&retmax to a big number to ensure I get all the tax_ids for that month (in this case 23511). I then made a local copy of the NCBI database in MySQL (
instructions here) and queried for all species-level taxa in GenBank. I used a rather crude regular expression
REGEXP '^[A-Z][a-z]+ [a-z][a-z]+$' to find just those species names that were likely to be proper scientific names (i.e., no "sp.", "aff.", museum or voucher codes, etc.). To group the species into major taxonomic groups I used the
division_id.
Results are available in a
Google Spreadsheet.









 My article describing
My article describing 
 What do crustaceans live on?
What do crustaceans live on?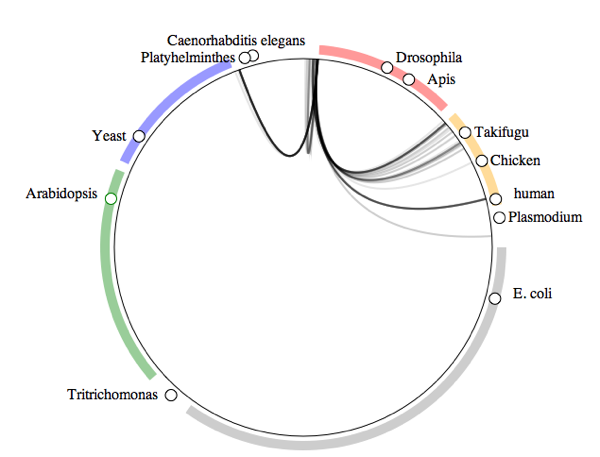

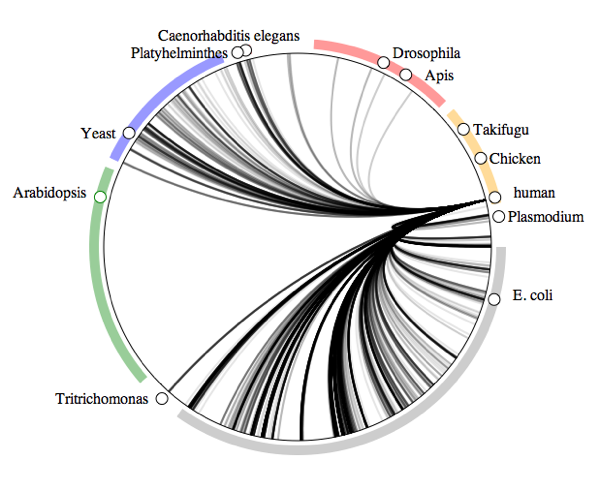



 How the visualisation was made
How the visualisation was made








 But what I've also noticed is that the site that I obtained the data from,
But what I've also noticed is that the site that I obtained the data from,