We invite participants for an event that will pioneer the mining of the Encyclopedia of Life (http://eol.org) and the Biodiversity Heritage Library (http://www.biodiversitylibrary.org) to address outstanding and novel questions about the ecology and evolution of biodiversity. We aim to identify questions and data for which biologists may lack informatics skills and resources to address or analyze successfully; and symmetrically, to guide informaticians to pressing ecological and evolutionary questions. We seek to make actual discoveries through joint activities and to test the “computability” of major biodiversity databases.Since I won't be applying to participate I thought I'd sketch some possible ideas here.
Co-occurrence of taxon names as proxy for ecological associations
Some time ago I noted that if you build a "tag tree" for taxonomic names in a BHL document you can get some interesting patterns, such as the names of hosts and their parasites occurring together. For example, searching BioNames for the rodent genus Praomys turns up papers with fleas, lice and cestodes. This suggests ways to mine BHL for ecological association data. It could be done by looking for general patterns of co-occurrence, or perhaps in a more targeted fashion (e.g., find all pages that have mammal and insect names together). Perhaps we could develop weighting schemes based on taxonomy whereby the co-occurrence of taxonomically unrelated groups is flagged as possibly significant (at the same time we'd want to avoid false positives such as tables of contents and indices).
Mining article titles for ecological associations
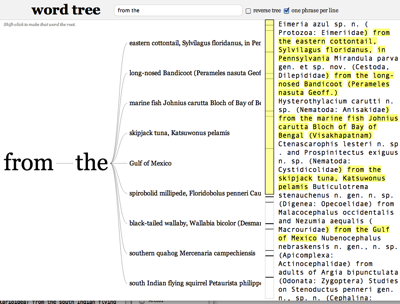
Another approach is to try and interpret the text itself. Keeping with the host-parasite theme, often descriptions of new parasite species are of the form "new species x from y". Here are some examples I use in my Phyloinformatics course:
Wordtrees are a great way to visualise these sentences and get insights into how to parse them (the word tree for the text above is here.
Geography
There is a lot of geographic data in BHL, which could potentially fill in gaps in geographic databases such as GBIF (which feeds into EOL). Even extracting latitude and longitude pairs from the OCR text can be enough to build some interesting maps.
Image extraction
Another approach is to extract images from BHL, ideally with the associated caption. This would be a way to quickly build an image database, a lot of taxonomic papers have illustrations of taxa, so this would be a quick way to get that information. It might be possible to do some clever parsing of the figure caption to extract not only taxon names but also other data. For example if the caption mentions a scale bar you could very quickly classify organisms into size categories (a 1mm scale bar versus a 1cm or 1m scale bar tells you something about the size of the organism).
Data extraction
Complementing the idea of image extraction, how about a tool that identifies tables in BHL OCR text? These tables are potentially sources of useful data, if they can be pulled out and indexed by taxon name (for example) then they could be analysed further. BHL OCR of tables tends to be poor, but the OCR could be redone on just the table, and/or the table could be edited manually (perhaps with the help of crowd sourcing).