The Global Names project will host a workshop to explore options and to make recommendations as to issues that relate to Attribution, Rights and Licensing of names and compilations of names. The aim of the workshop is a report that clarifies if and how we share names.
We seek submissions from all interested parties - nomenclaturalists, taxonomists, aggregators, and users of names. Let us know what (you think) intellectual property rights apply or what rights should be associated with names and compilations of names. How can those who compile names get useful attribution for names, and what responsibilities do they have to ensure that information is authoritative. If there are rights, what kind of licensing is appropriate.
Contributions can be submitted http://names-attribution-rights-and-licensing.wikia.com/wiki/Main_Page, where you will find more information about this event.
I'm trying to work out why this seemingly innocuous post made me so mad. I think this is because I think this fundamentally framing the question the wrong way. Surely the goal is to have a list of names that is global in scope, well documented, and freely usable by all without restriction? Surely we want open and free access to fundamental biodiversity data? In which case, can we please stop having meetings and get on with making this so?
If you frame the discussion as one of "Attribution, Rights and Licensing of names and compilations of names" then you've already lost sight of the prize. You've focussed on the presumed "rights" of name compilers instead.
I would argue that names compilations are somewhat overvalued. They are basically lists of names, sometimes (all to rarely) with some degree of provenance (e.g., a citation to the original use of the name). As I've documented before (e.g., More fictional taxa and the myth of the expert taxonomic database and Fictional taxa) entirely fictional can end up in taxonomic databases with alarming ease. So any claims that these are expert-curated lists should be taken with a pinch of salt.
Furthermore, it is increasingly easy to automate building these lists, given that we have tools for finding names in text, and an ever expanding volume of digitised text becoming available. Indeed, in an ideal world where all taxonomic literature was digitised much of the rationale for taxonomic name databases would disappear (in the same way that library card catalogues are irrelevant in the age of Google). We are fast approaching the point where we can do better than experts. To give just one example, in a recent BHL interview with Gary Poore it was stated that:
For example, the name widely used name Pentastomida itself was widely attributed to Diesing, 1836, but the word did not appear in the literature until 1905.
A quick check of Google Ngrams shows this to be simply false:
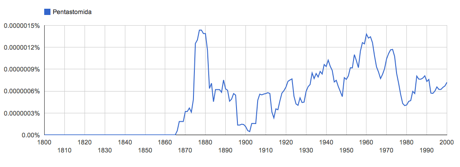
I don't need taxonomic expertise to see this, I simply need decent text indexing. So, if you have a list of names, you have something that it will soon be largely possible to recreate using automated methods (i.e., text mining). With a little sophistication we could mine the literature for further details, such as synonymy, etc. Annotation and clarification of a few "edge cases" where things get tricky will always be needed, but if you want to argue that your lists deserves "Attribution, Rights and Licensing" then you fail to realise that your list is going to be increasing easy to recreate simply by crawling the web.
It seems to me that most taxonomic databases are little more than digitised 5x3 index cards, and lack any details on the provenance of the names they contain. They often don't have links to the primary literature, and if they do cite that literature they typically do so in a way that makes it hard to find the actual publication. I once gave a talk which included the slide below showing taxonomic databases as being "in the way" between taxonomists and users of taxonomic information:

In the old days building taxonomic databases required expertise and access to obscure, hard to find, physical literature. A catalogue of names was a way to summarise that information (since we couldn't share access). Now we are in an age where more and more primary taxonomic information is available to all, which removes most of the rationale for taxonomic databases. Users can go directly to taxonomic information themselves, which mean they can get the "good stuff", and maybe even cite it (giving us provenance and credit, which I regard as basically the same thing). In many ways taxonomic databases are transitional phenomena (like phone directories, remember those), and one could argue are now in the way of the taxonomists' Holy Grail, getting their work cited.
Lastly, any discussion of "Attribution, Rights and Licensing of names and compilations of names" reflects one of the great self inflicted wounds of biodiversity informatics, namely the reluctance to freely share data. As we speak terrabytes of genomics data are whizzing around the planet, people are downloading entire copies of GenBank and creating new databases. All of this without people fussing over "Attribution, Rights and Licensing." It's time for taxonomic databases to get over themselves and focus on making biodiversity data as accessible and available as genomics data.