 Quick thoughts on the recent announcement by figshare and F1000 about the new journals being launched on the F1000 Research site. The articles being published have data sets embedded as figshare widgets in the body of the text, instead of being, say, a static table. For example, the article:
Quick thoughts on the recent announcement by figshare and F1000 about the new journals being launched on the F1000 Research site. The articles being published have data sets embedded as figshare widgets in the body of the text, instead of being, say, a static table. For example, the article:Oliver, G. (2012). Considerations for clinical read alignment and mutational profiling using next-generation sequencing. F1000 Research. doi:10.3410/f1000research.1-2.v1has a widget that looks like this:
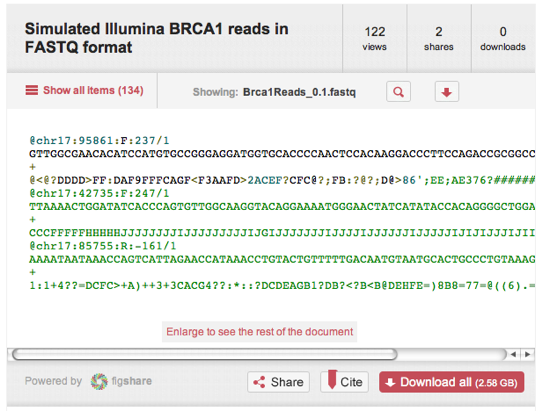
You can interact with this widget to view the data. Because the data are in figshare those data are independently citable, e.g. the dataset "Simulated Illumina BRCA1 reads in FASTQ format" has a DOI http://dx.doi.org/10.6084/m9.figshare.92338.
Now, wouldn't it be cool if TreeBASE did something similar? Imagine if uploading trees to TreeBASE were easy, and that you didn't have to have published yet, you just wanted to store the trees and make them citable. Imagine if TreeBASE had a nice tree viewer (no, not a Java applet, a nice viewer that uses SVG, for exmaple). Imagine if you could embed that tree viewer as a widget when you published your results. It's a win all round. People have an incentive to upload trees (nice viewer, place to store them, and others can cite the trees because they'd have DOIs). TreeBASE builds its database a lot more quickly (make it dead easy to upload tree), and then as more publishers adopt this style of publishing TreeBASE is well placed to provide nice visualisations of phylogenies pre-packaged, interactive, and citable. And let's not stop there, how about a nice alignment viewer? Perhaps this is the something currently rather moribund PLoS Currents Tree of Life could think about supporting?