Recently I’ve been exploring data downloaded from BOLD. Part of this was motivated by work done with David Schindel for a recent book:
Schindel, D.E., Page, R.M.P. (2024). Creating Virtuous Cycles for DNA Barcoding: A Case Study in Science Innovation, Entrepreneurship, and Diplomacy. In: DeSalle, R. (eds) DNA Barcoding. Methods in Molecular Biology, vol 2744. Humana, New York, NY. doi:10.1007/978-1-0716-3581-0_1
In this blog post I record some struggles I’ve had with the supposedly “Frictionless” data provided by BOLD. I list a serious of issues, and make some recommendations as to how these can be fixed.
Previous versions disappear from site
The web page Data Packages lists datasets that can be downloaded.
The two most recent have the DOIs:
While this makes it easy to link to the latest version of the data, it inhibits reproducibility because the data doi:10.5883/DP-Latest points to can change, and there is (currently) no unique DOI for that particular dataset. Once a version becomes the third oldest, then it seems to get a version-specific DOI.
The list of “Historical Data” releases is not exhaustive. Currently (2024-10-17) there are four older versions listed:
However I have downloaded older versions that are no longer listed on the Data Packages web page. This makes it hard for anyone wanting to trace the history of changes in BOLD data.
Recommendation
Have distinct DOIs for latest version (i.e., in addition to “10.5883/DP-Latest”), and keep list of all releases on the web site.
Even better, switch to using Zenodo to store data, they provide a nicer model of versioning, and can also provide download metrics.
Where are the images?
A major surprise is the lack of URLs for specimen images. The imagery in BOLD is very useful, yet not included in the data export! The only way to get a list of image URLs is to get the data from GBIF(!).
Recommendation
Include images URLs in the data releases.
Column names change over time and aren’t standardised
One major source of frustration is that the labels used for the columns of data can change. This is, pun intended, a major source of friction in supposedly “frictionless” data. Code that successfully parses one dataset may fail with the new release. One could argue that the code should rely solely on the information in the data package, but there are some columns (e.g., geographic coordinates, the raw sequences) that require special treatment, and the natural language descriptions of each column are not machine-readable.
Below is a table of column names from a range of data packages from 2022-09-28 to 2024-09-06. Most column names are stable, but sometimes new ones appear, and sometimes they vanish. Core data elements, such as nucleotide sequences can change, e.g. nuc versus nucraw.
| column name | 2022-09-28 | 2023-09-29 | 2023-10-27 | 2024-07-26 | 2024-08-02 | 2024-08-09 | 2024-09-06 |
|---|
| associated_specimen | | | | | | | |
| associated_specimens | | | | | | | |
| associated_taxa | | | | | | | |
| bin_created_date | | | | | | | |
| bin_uri | | | | | | | |
| biome | | | | | | | |
| bold_recordset_code_arr | | | | | | | |
| class | | | | | | | |
| collection_code | | | | | | | |
| collection_date | | | | | | | |
| collection_date_accuracy | | | | | | | |
| collection_date_end | | | | | | | |
| collection_date_start | | | | | | | |
| collection_event_id | | | | | | | |
| collection_note | | | | | | | |
| collection_notes | | | | | | | |
| collection_time | | | | | | | |
| collectors | | | | | | | |
| coord | | | | | | | |
| coord_accuracy | | | | | | | |
| coord_source | | | | | | | |
| country | | | | | | | |
| country/ocean | | | | | | | |
| country_iso | | | | | | | |
| depth | | | | | | | |
| depth_accuracy | | | | | | | |
| ecoregion | | | | | | | |
| elev | | | | | | | |
| elev_accuracy | | | | | | | |
| extrainfo | | | | | | | |
| family | | | | | | | |
| fieldid | | | | | | | |
| funding_src | | | | | | | |
| gb_acs | | | | | | | |
| genus | | | | | | | |
| geoid | | | | | | | |
| habitat | | | | | | | |
| identification | | | | | | | |
| identification_method | | | | | | | |
| identification_rank | | | | | | | |
| identified_by | | | | | | | |
| identifier_email | | | | | | | |
| insdc_acs | | | | | | | |
| inst | | | | | | | |
| kingdom | | | | | | | |
| life_stage | | | | | | | |
| marker_code | | | | | | | |
| museumid | | | | | | | |
| notes | | | | | | | |
| nuc | | | | | | | |
| nuc_basecount | | | | | | | |
| nucraw | | | | | | | |
| order | | | | | | | |
| phylum | | | | | | | |
| primers_forward | | | | | | | |
| primers_reverse | | | | | | | |
| processid | | | | | | | |
| processid_minted_date | | | | | | | |
| province | | | | | | | |
| realm | | | | | | | |
| record_id | | | | | | | |
| recordset_code_arr | | | | | | | |
| region | | | | | | | |
| reproduction | | | | | | | |
| sampleid | | | | | | | |
| sampling_protocol | | | | | | | |
| sector | | | | | | | |
| sequence_run_site | | | | | | | |
| sequence_upload_date | | | | | | | |
| sex | | | | | | | |
| short_note | | | | | | | |
| site | | | | | | | |
| site_code | | | | | | | |
| sovereign_inst | | | | | | | |
| species | | | | | | | |
| species_reference | | | | | | | |
| specimen_linkout | | | | | | | |
| specimenid | | | | | | | |
| subfamily | | | | | | | |
| subspecies | | | | | | | |
| taxid | | | | | | | |
| taxon_name | | | | | | | |
| taxon_rank | | | | | | | |
| taxonomy_notes | | | | | | | |
| tissue_type | | | | | | | |
| tribe | | | | | | | |
| voucher_type | | | | | | | |
Recomendation
Avoid changing the names of data columns between releases. Adopt standardised terms, such as Darwin Core, wherever possible. Tell us that these are Darwin Core by using the dwc: prefix.
Use identifiers for people
People appear in several places in the data, notably as identifiers and collectors. The BOLD data uses simple text strings (i.e., names) of people, rather than external identifiers such as ORCIDs. This means we miss out on valuable information. For example, for a 2 million sequence subset of the latest release I was curious as to who identified the most specimens. For each of these names I then wanted to find out who they were. For example, are they taxonomists? If so, what is their expertise? What taxonomic papers have they published? Where are they based? If BOLD included ORCID ids it would be easier to answer these questions. Instead, I resorted to Google and produced the following table:
Top 10 identifiers of BOLD specimens:
Note that most of the top ten identifiers work at the University of Guelph, the home of BOLD. This tells us something about the degree to which BOLD is dependent on its own staff to identify specimens, versus the extent to which it has engaged the wider community.
The flip side of this is that these people are curating an important database. Are they getting credit? Is this curation making its way into Bionomia, which has mechanisms to give credit to this work.
I have also briefly looked at collector names, and it is - as one might expect - something of a mess. The same person’s name is written different ways, text strings representing multiple people are incorrectly split into individual names, etc. The description in the data package is more wishful thinking than reality:
Comma separated list of full or abbreviated names of the individuals or teams responsible for collecting the sample in the field.
Recommendation
Add ORCID identifiers for people who have identified specimens.
Method of identification not standardised
The value of BOLD as a tool for identifying new sequences depends on the reliability of existing DNA barcodes. How are these identified? The field identification_method is full of a mix of terms. There are all the obvious traps people fall into when not being careful with data. The same term may be spelt differently and/or is capitalised differently. People add qualifiers to a term, such as the date of identification, making it much harder to ask simple questions such as how many sequences have been identified based on their morphology, versus based on their sequences.
So far I’ve found 1889 different terms for identification method, here are the top 20:
- BIN Taxonomy Match
- BOLD ID Engine Manual
- BOLD Sequence Classifier
- Morphology
- morphology
- morphological
- BOLD ID Engine (March 2015)
- Tree based Identification(April 2016)
- Morphological
- BIN Taxonomy Match (Mar 2023)
- BIN Taxonomy Match (May 2019)
- Tree based Identification (Feb 2017)
- BIN Taxonomy Match (May 2017)
- BIN Taxonomy Match (Oct 2022)
- BIN Taxonomy Match (Aug 2023)
- BIN Taxonomy Match (Mar 2017)
- BOLD ID Engine
- BIN Taxonomy Match (Apr 2017)
- BIN Taxonomy Match (Jun 2019)
- Tree Based Identification (April 2016)
Note that we have “Morphology”, “morphology”, “morphological”, and “Morphological”. How is “BOLD ID Engine Manual” different from “BOLD ID Engine”? Note the use of dates as qualifiers.
Recommendation
Enforce a standardised vocabulary, add additional fields for date and notes on identification.
Voucher type not standardised
The description of the voucher_type reads:
“Status of the specimen in an accessioning process.This field uses a controlled vocabulary: ‘Museum Vouchered:Type’, ‘Museum Vouchered:Type Series’, ‘Vouchered:Registered Collection’, ‘To Be Vouchered:Holdup/Private’, ‘E-Vouchered:DNA/Tissue+Photo’, ‘Dna/Tissue Vouchered Only’, ‘No Specimen’.”
This is patently false. Instead of seven terms are at least 508 for this field. Here are the top 20 terms (* indicates a term from the controlled vocabulary):
- Vouchered:Registered Collection*
- DNA/Tissue Vouchered Only*
- To Be Vouchered:Holdup/Private*
- museum voucher
- E-Vouchered:DNA/Tissue+Photo*
- Voucher Type: Morphological
- No Specimen*
- Museum voucher, whole specimen in ethanol
- Vouchered:Private Collection
- Museum Vouchered:Type*
- Museum Vouchered:Type Series*
- Museum voucher, Whole specimen in ethanol
- Museum voucher, whole specimen
- Museum Vouchered
- Museum voucher, e-voucher
- Museum voucher, Whole specimen
- Museum voucher, E-vouchered with additional representatives stored in ethanol in parent lot
- vouchered: not registered collection
- Museum voucher, E-vouchered with additional representatives stored in ethanol
- in alcohol (ethanol, 96%)
Recommendation
Enforce the existing controlled vocabulary.
Institutions lack identifiers
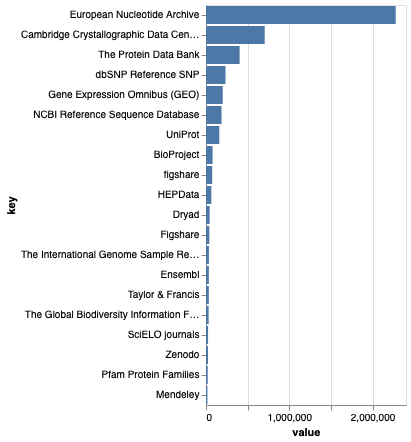
Institutions are listed by name. As with any string, there is the potential for different spellings and formatting. Anyone interested in getting metrics for institutional engagement with BOLD, and comparing that to, say, sources of funding, would much rather have identifiers than strings.
Here are the top 20 institutions:
- Centre for Biodiversity Genomics
- University of Pennsylvania
- Area de Conservacion Guanacaste
- Mined from GenBank, NCBI
- Canadian National Collection of Insects, Arachnids and Nematodes
- SNSB, Zoologische Staatssammlung Muenchen
- Australian National Insect Collection
- University of Malaya, Museum of Zoology
- Instituto Nacional de Biodiversidad, Costa Rica
- Royal Ontario Museum
- California Academy of Sciences
- NEON Biorepository at Arizona State University
- Research Collection of M. Alex Smith
- Smithsonian Tropical Research Institute
- University of New Brunswick, Fredericton
- York University, Packer Collection
- Wellcome Sanger Institute
- Smithsonian Institution, National Museum of Natural History
- Natural History Museum, London
- University of Oulu, Zoological Museum
Recommendation
Add external identifiers for institutions such as RORs.
Summary
Some of these issues raised here are easy to fix, other will require a lot of curation. I suspect that part of the problem is that there’s no evidence that BOLD itself makes use of these data dumps. If you view data exports as somethign you are “supposed to” do, rather than something that you yourself use, then there’s no incentive to make sure the data is fit for purpose. Eating your own dog food is a great way to avoid these problems.
Written with StackEdit.