 In a classic paper Boggs (1949) appealed for an “atlas of ignorance”, an honest assessment of what we know we don’t know:
In a classic paper Boggs (1949) appealed for an “atlas of ignorance”, an honest assessment of what we know we don’t know:
Boggs, S. W.. (1949). An Atlas of Ignorance: A Needed Stimulus to Honest Thinking and Hard Work. Proceedings of the American Philosophical Society, 93(3), 253–258. Retrieved from http://www.jstor.org/stable/3143475
This is the theme of this year's GBIF Challenge: Analysing and addressing gaps and biases in primary biodiversity data. "Gaps" can be gaps in geographic coverage, taxa group, or types of data. GBIF is looking for ways to access the nature of the gaps in the data it is aggregating from its network of contributors.
How to enter
Details on how to enter are on the Challenge website, deadline is September 30th.Ideas
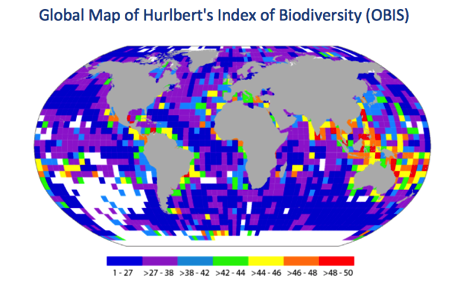
One approach to gap analysis is to compare what we expect to see with what we actually have. For example, we might take a “well-known” group of organisms and use that to benchmark GBIF’s data coverage. A drawback is that the “well-known” organisms tend to be the usual suspects (birds, mammals, fish, etc.), and there is the issue of whether the chosen group is a useful proxy for other taxa. Another approach is to base the estimate of ignorance on the data itself. For example, OBIS has computed Hurlbert's index of biodiversity for its database, e.g. http://data.unep-wcmc.org/datasets/16
 Can we scale these methods to the 600+ million records in GBIF? There are some clever methods for using resampling methods (such as the bootstrap) on large data sets that might be relevant, see http://www.unofficialgoogledatascience.com/2015/08/an-introduction-to-poisson-bootstrap_26.html.
Can we scale these methods to the 600+ million records in GBIF? There are some clever methods for using resampling methods (such as the bootstrap) on large data sets that might be relevant, see http://www.unofficialgoogledatascience.com/2015/08/an-introduction-to-poisson-bootstrap_26.html.
Another approach might be to compare different datasets for the same taxa, particularly if one data set is not in GBIF. Or perhaps we can compare datasets for the same taxa collected by different methods.
Or we could look at taxonomic gaps. In an earlier post The Zika virus, GBIF, and the missing mosquitoes I noted that GBIF's coverage of vectors of the Zika virus was very poor. How well does GBIF cover vectors and other organisms relevant to human health? Maybe we could generalise this to explore other taxa. It might, for example, be interesting to compare degree of coverage for a species with some measure of the "importance" of that species. Measures of importance could be based on, say, number of hits in Google Scholar for that species, size of Wikipedia page (see Wikipedia mammals and the power law), etc.
Gaps might also be gaps in data completeness, quality, or type.