For simplicity let's suppose that we have this basic model:
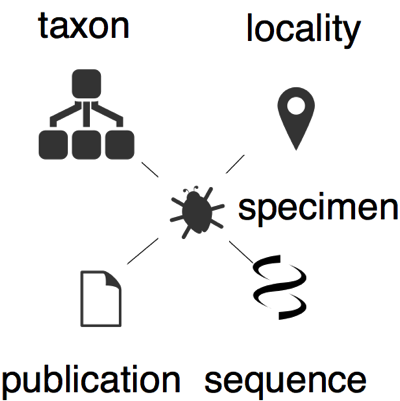
A specimen comes from a locality (ideally we have the latitude and longitude of that locality), it is assigned to a taxon, we have data derived from that specimen (e.g., one or more DNA sequences), and we have one or more publications about that specimen (e.g., a paper that publishes a taxon name for which the specimen is a type, or a paper that publishes a sequence for which the specimen is a voucher).
NCBI
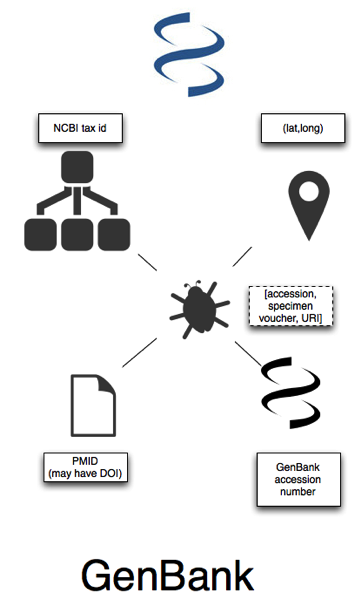
In GenBank we have sequences that have accession numbers, and these are linked to taxa (identified by NCBI tax ids). A nice feature of sequence databases is that taxa are explicitly defined by extension, that is, a taxon is the set of sequences assigned to a given taxon. Most (but not all, see Miller et al. doi:10.1186/1756-0500-2-101) sequences are also linked to a publication, which will usually have a PubMed id (PMID), and sometimes a DOI. Many sequences are also georeferenced (see Guest post: response to "Putting GenBank Data on the Map"). Most sequences aren't linked to a voucher specimen, but there is the implict notion of a source (in RDF-speak, many specimens are "blank nodes" Blank nodes for specimens without URI). Some sequences are associated with a specimen that has a museum code, and some are explicitly linked to the specimen by a URL.
DNA barcodes
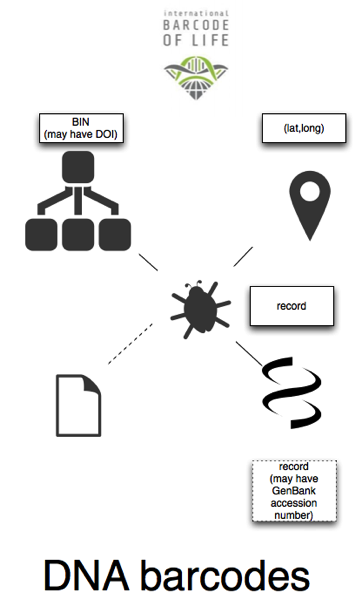
Barcodes, as represented in BOLD are similar to sequences in GenBank. We have explicit taxa ("BINs") each of which has a URL, some also having DOIs. Most barcodes are georeferenced. There's some ambiguity about whether the URL for a barcode record identifies the barcode sequence, the specimen, or both. There may be a voucher code for the specimen. Some barcodes are linked to publications, but not (as far as I can see) in the data obtained from the API. Some barcodes are linked to the corresponding record in GenBank (which may or may not be supressed, see Dark taxa even darker: NCBI pulls (some) DNA barcodes from GenBank (updated)).
GBIF
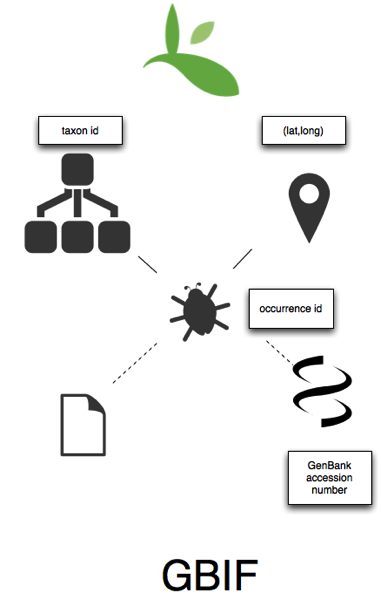
At it's core GBIF has occurrence records (many of these are specimen-based, but the majority of data in GBIF is actually observation-based), each of which has a unique id, and which is linked to a taxon, also with a unique id. As with the sequence databases, a taxon is a set of occurrences that have been assigned to that taxon. Many records in GBIF are georeferenced. There are limited cross links to other database - some occurrences list associated GenBank sequences. Some GBIF occurrences actually are sequences (e.g., the European Molecular Biology Laboratory Australian Mirror and the soon to be indexed Geographically tagged INSDC sequences), and barcodes are also making their way into GBIF (e.g., Zoologische Staatssammlung Muenchen - International Barcode of Life (iBOL) - Barcode of Life Project Specimen Data). Links to publications are limited.
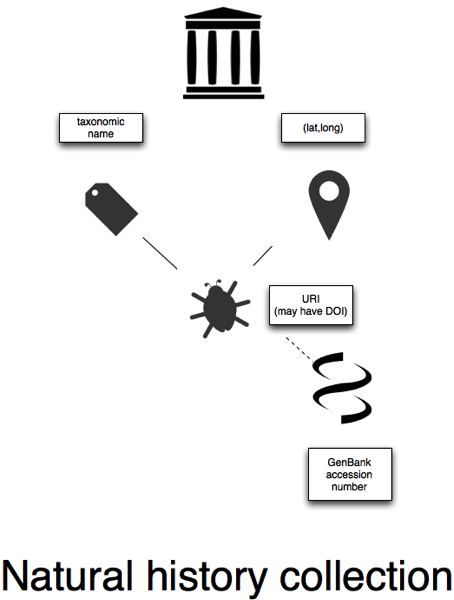
Museums and herbaria
Some individual natural history collections which are online provide specimen-level web pages and URLs (some even have DOIs, see DOIs for specimens are here, but we're not quite there yet), and some museums list associated GenBank sequences. In the diagram I've not linked the specimens to a taxon, because most specimens are tagged by a name, not an explicit taxon concept (unlike NCBI, BOLD, or GBIF).
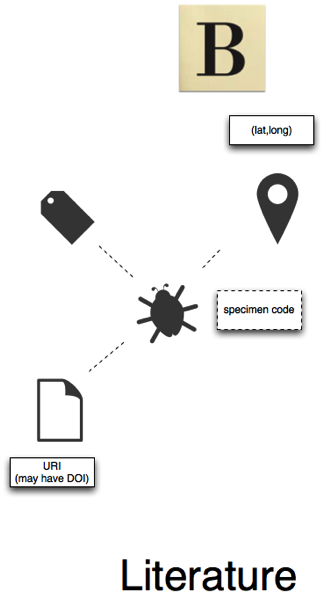
Literature
Literature databases (represented here by BioStor, but could be other sources such as ZooKeys, for example) may contain articles that mention specimen codes. These articles may also mention taxon names, and geographic localities (including coordinates) (see, for example, Linking GBIF and the Biodiversity Heritage Library. Mining text for names, specimens, and localities is fairly easy, but linking these together is harder (i.e., this specimen is of this taxon, and was found at this locality).
Linking together
If we have these separate sources and this trivial model, then we can imagine trying to tie information about the same specimen together across the different databases. Why might we want to do this. Here are three reasons:
- Augmentation Combining information can enhance our understanding of a specimen. Perhaps a specimen in GBIF is a geographic outlier. A publication that mentions the specimen includes it in a new taxon, perhaps discovered by sequencing DNA extarcted from that specimen. Linking this information together resolves the problematic distribution.
- Provenance What is the evidence that a particular specimen belongs to a particualr taxon, or was collected at a particular locality? If we connect specimens to the literature we we can review the evidence for ourselves. If we have sequences we can run BLAST, build a tree, and see if we should rethink our classification of that sequence. Imagine being able to browse GBIF and see the evidence for each dot on the map?
- Citation Mentions in the literature, use as vouchers for DNA barcoding or other forms of sequencing can be thought of a "citation" of that specimen. Museums hosting that material could use metrics base don this to demonstrate the value of their collection (see also The impact of museum collections: one collection ≈ one Nobel Prize).
Making the links
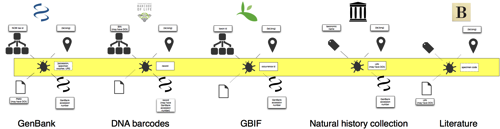
All this is well and good, the trick is to actually make the links. Here things get horribly messy very quickly. Museum specimens are cited in inconsistent ways, we don't have widely used unique, resolvable specimen identifiers, and even if we did have these identifiers we don't have a global discovery mechanism for matching voucher codes to identifiers. GBIF would be an obvious part of a "global discovery mechanism" (bit like CrossRef but for specimens), GBIF can have multiple records for the same specimen. Sometimes this is because GBIF not only aggregates data from primary sources (such as museums) but also other aggregations which may themselves already include specimens harvested from primary sources. GBIF can also have multiple records because museums keep messing with their databases, try new variants of the Darwin Core triple, etc., resulting in records that look "new" to GBIF. Whole collections can be duplicate din this way.
One way to tackle this multiplicity of specimen records is to think in terms of "clusters" of specimens that are, in some sense, the same thing across multiple databases. For example, clustering a set of duplicated GBIF records together with the sequences derived from those specimens, perhaps including a DNA barcode, and a list of papers that mention that specimen. This is represented by the yellow bar through the diagram, it connects all the different pieces of information about a specimen into a single cluster. More *cough* later.