Gastrotheca aureomaculata Cochran and Goin, 1970, Bull. U.S. Natl. Mus., 288: 177. Holotype: FMNH 69701, by original designation. Type locality: "in [Departamento] Huila, Colombia, at San Antonio, a small village 25 kilometers west of San Agustín, at 2,300 meters".
The journal that ASW abbreviates as "Bull. U.S. Natl. Mus." is in the BHL, which gives its title as "Bulletin - United States National Museum.". How do I link these two records? In my bioGUID OpenURL project I've been doing things like using SQL LIKE statements with periods (.) replaced by wildcards ('%') to find journal titles that match abbreviations (as well as building a database of these abbreviations). But this is error prone, and won't work for abbreviations such as "Bull. U.S. Natl. Mus." because the word "National" has been abbreviated to "Natl", which isn't a substring of "National".
After exploring various methods (including longest common subsequences, and sequence alignment algorithms) I came across a MySQL plugin for n-grams. The plugin tokenises strings into bi-grams (tokens with just two characters, see the Wikipedia page on N-grams for more information). This means that even though as words "National" and "Natl" are different, they will have some similarity due to the shared bi-grams "Na" and "at".
So, I grabbed the source for the plugin and the ICU dependency, compiled the plugin and added it to MySQL (I'm running MySQL 5.1.34 on Mac OS X 10.5.8). The plugin can be added while the MySQL server is running using this SQL command:
INSTALL PLUGIN bigram SONAME 'libftbigram.so';
Initial experiments seem promising. For the bhl_title table I created a bi-gram index:
ALTER TABLE `bhl_title` ADD FULLTEXT (`ShortTitle`) WITH PARSER bigram;
If I then take the abbreviation "Bull. U.S. Natl. Mus.", strip out the punctuation, and search for the resulting string ("Bull US Natl Mus")
SELECT TitleID, ShortTitle, MATCH(ShortTitle) AGAINST('Bull U S Natl Museum')
AS score FROM bhl_title
WHERE MATCH(ShortTitle) AGAINST('Bull U S Natl Museum') LIMIT 5;
I get this:
| TitleID | ShortTitle | score |
|---|---|---|
| 7548 | Bulletin - United States National Museum. | 19.4019603729248 |
| 13855 | Bulletin du Muséum National d'Histoire Naturelle. | 17.6493873596191 |
| 14964 | Bulletin du Muséum National d'Histoire Naturelle. | 17.6493873596191 |
| 5943 | Bulletin du Muséum national d'histoire naturelle. | 17.6493873596191 |
| 12908 | Bulletin du Muséum National d'Histoire Naturelle. | 17.6493873596191 |
The journal we want is the top hit (if only just). I'll probably have to do some post-processing to check that the top hit makes sense (e.g., is it a supersequence of the search term?) but this looks like a promising way to match abbreviated journal names and book titles to records in BHL (and other databases).
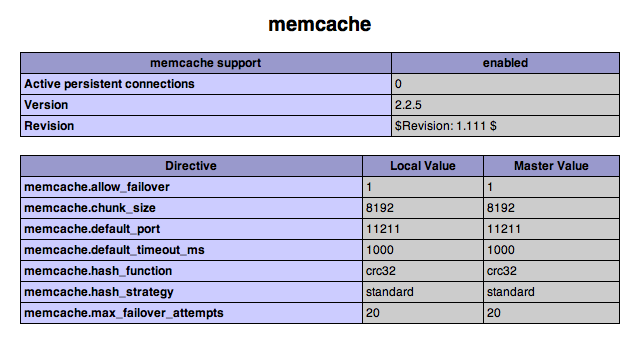

 One thing I find myself doing (probably more often than I should) is adding a reference to
One thing I find myself doing (probably more often than I should) is adding a reference to  The
The  Wikipedia illustrates this nicely. Wikipedia conforms to Gregg's model in that taxa are defined by extension (each taxon comprises one or more wiki pages), and if taxa have the same content only one taxon (typically that with the lowest taxonomic rank) has a page in Wikipedia. Put another way, if the aardvark is the sole representative of the Tubulidentata, then there is nothing that could be put on the Tubulidentata page that shouldn't also belong on the page for the aardvark. As a result, the page for the
Wikipedia illustrates this nicely. Wikipedia conforms to Gregg's model in that taxa are defined by extension (each taxon comprises one or more wiki pages), and if taxa have the same content only one taxon (typically that with the lowest taxonomic rank) has a page in Wikipedia. Put another way, if the aardvark is the sole representative of the Tubulidentata, then there is nothing that could be put on the Tubulidentata page that shouldn't also belong on the page for the aardvark. As a result, the page for the